Difference between revisions of "Normal Mode Analysis Hemochromatosis"
Bernhoferm (talk | contribs) (→Modes) |
Bernhoferm (talk | contribs) (→Modes) |
||
| Line 187: | Line 187: | ||
| align="right" | [[File:hemo_elnemo_mode12_front.gif|thumb|150px|Mode 12 (front)]] |
| align="right" | [[File:hemo_elnemo_mode12_front.gif|thumb|150px|Mode 12 (front)]] |
||
|- |
|- |
||
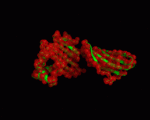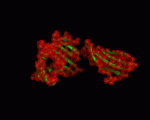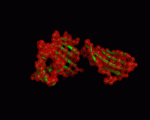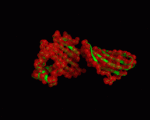
| − | |+ style="caption-side: bottom; text-align: left" |<font size=1>'''Table X:''' Visualization of the normal modes 7 to 12 for 1a6zC (by ElNemo). The base structure of HFE is shown in green, the normal modes in red |
+ | |+ style="caption-side: bottom; text-align: left" |<font size=1>'''Table X:''' Visualization of the normal modes 7 to 12 for 1a6zC (by ElNemo). The base structure of HFE is shown in green, the normal modes in red. Each mode is shown from the side and front of the protein. |
|} |
|} |
||
</figtable> |
</figtable> |
||
Revision as of 22:19, 27 July 2012
Hemochromatosis>>Task 9: Normal mode analysis
Contents
Short task description
Detailed description: Normal mode analysis
Protocol
A protocol with a description of the data acquisition and other scripts used for this task is available here.
General
<figtable id="comparison">
</figtable>
We used the PDB structures 1a6z and 1de4 for our NMA. 1a6z represents HFE bound with Beta-2-Microglobulin only and 1de4 contains the whole TFR-HFE-B2M complex. We extracted HFE's conformations from these structures (chain C from 1a6z and chain A from 1de4). Both files contain the same 272 residues (26-297 within HFE's sequence). A comparison between the two showed that there are only minor differences in the conformation (cf. <xr id="comparison"/>) and almost all of them are within loop regions. TM-Score calculated an RMSD of only 1.602 and a TM-Score of 0.9607 for them.
For the NMA we used WEBnm@ and ElNemo. Both webservers use the C-alpha atoms only for the normal mode calculations and thus could theoretically compute 816 (3N, N = 272) normal modes, though the first 6 are irrelevant as they represent the three simple translations and rotations of the whole protein. In addition to the normal modes they provide also features to compare normal mode motions with other conformations of the same protein or correlation analysis of different normal modes.
WEBnm@
The deformation energy scores computed by WEBnm@ are shown in <xr id="webnma_energy"/>. The first 6 modes (7-12) could be put into three groups: Mode 7-9 which have a very small score, 10 and 11 which have about double the score of 8/9, and Mode 12 which again doubles Mode 10's score. This suggests that the first three combine big motions while the latter three exhibit more and more smaller parts that move separately.
<figtable id="webnma_energy">
| Mode number | Energy score | Mode number | Energy score |
|---|---|---|---|
| 7 | 416.65 | 14 | 4673.81 |
| 8 | 841.18 | 15 | 5862.82 |
| 9 | 849.58 | 16 | 6800.91 |
| 10 | 1436.11 | 17 | 8236.15 |
| 11 | 1899.61 | 18 | 7878.34 |
| 12 | 2981.09 | 19 | 8104.36 |
| 13 | 4518.38 | 20 | 9899.01 |
</figtable>
Modes
<figtable id="webnma_modes7to12">
</figtable>
Correlation of motions
<figure id="correlation">
</figure>
<xr id="correlation"/> shows the correlated motions of all residues within 1a6zC based on the calculated normal modes. At first sight the matrix could be separated into two areas: 1-180 and 181-272. The first one could further be separated into 1-90 and 91-180. When adjusted for the missing residues (1-25) in the PDB file the regions would correspond to 26-115, 116-205, and 206-297 respectively. According to Uniprot these numbers match the residues for HFE's alpha 1 (23-114), alpha 2 (115-205), and alpha 3 (206-297) domains. Though the differentiation in the 1-180 area is not that simple if you are not looking for three domains.
Overlap
<figtable id="overlap">
</figtable>
Atomic fluctuations
<figtable id="webnma_fluctuations">
</figtable>
ElNemo
Modes
<figtable id="elnemo_modes7to12">
</figtable>
Overlap
References
<references/>