Phenylketonuria
Phenylketonuria (PKU) is an inborn metabolic disorder and is caused by different mutations in the enzyme phenylalanine hydroxylase (PAH), which is responsible for the alteration of phenylalanine to tyrosine. Untreated PKU can lead to psychotic disorders and the postnatal development of the brain can be harmed.
Contents
Phenotype
PKU results in enrichment of phenylalanine and decrease of tyrosine, whereby other features of the phenylalanine metabolism pathway are affected like the production of melanin. Thus symptoms like fair skin, light hair and blue or sometimes even red eyes can occur. Additionally there can be mousy odour, eczema, epilepsy and perceptible posture changes. In some cases the concerned persons show psychotic disorders, ADHD (attention-deficit hyperactivity disorder), Aggressions, autistic features, Hyperactivity and Self-harm (Self-mutilation).
There are two different PKU variants defined:
- Classical Phenylketonuria is caused by mutation/s in the PAH gene (precisely described here)
- Atypical (mild) Phenylketonuria is a deficiency of the tetrahydrobiopterin (BH4), a cofactor of the enzyme phenylalanine hydroxylase
Cross-references
See also description of this disease in
Treatment
After birth every child is tested with a newborn-screening, where some disorders are analysed. If a higher concentration of phenylalanine or a bad ratio of phenylalanine and tyrosine is found, PKU is diagnosed and the patient can be treated as followed:
- Special low phenylalanine diet for the whole life (routine treatment; constrains in meat, fish, eggs, nuts, legumes and milk products and careful usage in starchy food)
- Maternal PKU: If women with phenylketonuria wish to have a child, they have to do a strict diet before and during pregnancy.
- Tyrosine enriched food or medicaments
- Intake of the enzyme PAL (phenylalanine ammonia lyase), which converts phenylalanine to a harmless metabolite
- Tetrahydrobiopterin (only for mild PKU)
Cross-references
Biochemical disease mechanism
<figure id="pku_path">
</figure>
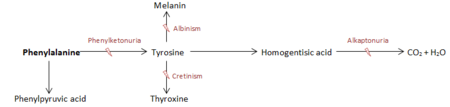
- The symptoms of PKU are caused by the complete or partial defected catalysing of phenylalanine to tyrosine.
- As it occurs at the beginning of the phenylalanine metabolism pathway other metabolic diseases are associated like Albinism or Cretinism (<xr id="pku_path"/>)
- PKU was named after the phenylketone (phenylpyruvate) in which phenylalanine is converted.
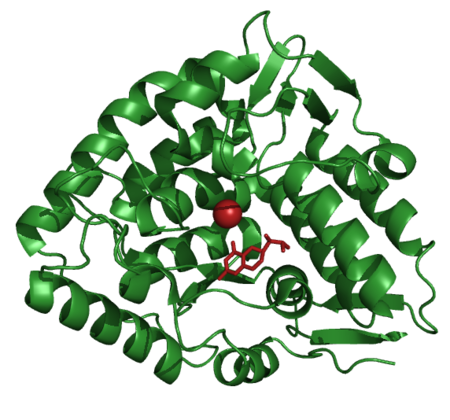
- Protein phenylalanine hydroxylase:
- is expressed in liver,
- has a sequence length of 452 aa,
- is encoded by the gene PAH (see Genomic information) on chromosome 12.
<figure id="metPath">

</figure>
Mutations and Genomic information
Genomic information
<figure id="chr12">
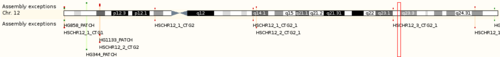
</figure>
PKU is an autosomal recessive disorder which can be caused by 400 different known mutations on the PAH gene. This gene lies on the human chromosome 12 (<xr id="chr12"/>) and consists of 13 exons. Most common are missense mutations, but also nonsense and silent mutations can be found. Normally the enzyme phenylalanine hydroxylase converts phenylalanine into tyrosine. If this pathway is disturbed, it is transformed to phenylpyruvic acid which constrains pyruvate decarboxylase in the brain (<xr id="hydrox"/>). This seems to be the reason for the mental disorders.
<figure id="hydrox">
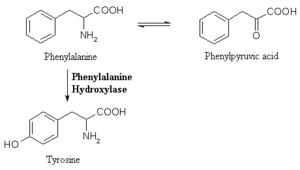
</figure>
Reference sequence
The Uniprot entry for PAH P00439 gives the protein sequence used as reference in this practical:
>sp|P00439|PH4H_HUMAN Phenylalanine-4-hydroxylase OS=Homo sapiens GN=PAH PE=1 SV=1 MSTAVLENPGLGRKLSDFGQETSYIEDNCNQNGAISLIFSLKEEVGALAKVLRLFEENDV NLTHIESRPSRLKKDEYEFFTHLDKRSLPALTNIIKILRHDIGATVHELSRDKKKDTVPW FPRTIQELDRFANQILSYGAELDADHPGFKDPVYRARRKQFADIAYNYRHGQPIPRVEYM EEEKKTWGTVFKTLKSLYKTHACYEYNHIFPLLEKYCGFHEDNIPQLEDVSQFLQTCTGF RLRPVAGLLSSRDFLGGLAFRVFHCTQYIRHGSKPMYTPEPDICHELLGHVPLFSDRSFA QFSQEIGLASLGAPDEYIEKLATIYWFTVEFGLCKQGDSIKAYGAGLLSSFGELQYCLSE KPKLLPLELEKTAIQNYTVTEFQPLYYVAESFNDAKEKVRNFAATIPRPFSVRYDPYTQR IEVLDNTQQLKILADSINSEIGILCSALQKI
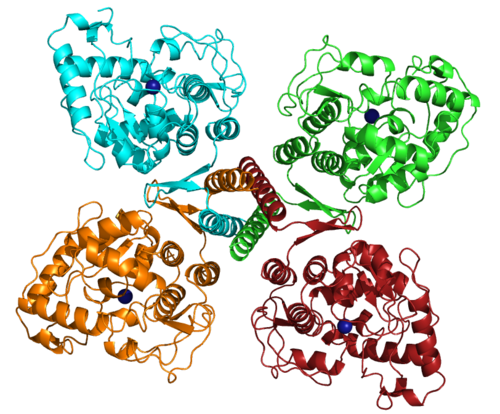
3D structure
</figure> </figure>| <figure id="1j8u"> |
| <figure id="2pah"> |
Database identifiers
In <xr id="databases"/> most databases containing information about PAH or PKU itself used for this practical can be viewed. For the structures found in RSCB there are a lot of different entries concerning PAH. The two IDs represent the structures used by us. <figtable id="databases">
| Database | PAH-ID |
|---|---|
| UniProt | P00439 |
| RSCB | 2PAH, 1J8U* |
| Ensembl | ENSG00000171759 |
| OMIM | 612349 (PAH), 261600 (PKU) |
| dbSNP | PAH, PKU |
</figtable>
Tasks
Task 2: Sequence searches and multiple sequence alignments
Task 3: Sequence-based predictions
Task 4: Structural Alignments
Task 5: Homology based structure prediction
Task 6: Protein structure prediction
Task 7: Researching SNPs
Task 8: Sequence-based mutation analysis
Task 9: Structure-based mutation analysis
Task 10: Normal mode analysis
References
<references/>