Gaucher Disease: Task 04 - Structural Alignment
<css>
table.colBasic2 { margin-left: auto; margin-right: auto; border: 1px solid black; border-collapse:collapse; }
.colBasic2 th,td { padding: 3px; border: 1px solid black; }
.colBasic2 td { text-align:left; }
/* for orange try #ff7f00 and #ffaa56 for blue try #005fbf and #aad4ff
maria's style blue: #adceff grey: #efefef
- /
.colBasic2 tr th { background-color:#efefef; color: black;} .colBasic2 tr:first-child th { background-color:#adceff; color:black;}
</css>
This page is still under construction.
Contents
Exploring Structural Alignments
Used data set
<figtable id="data_set">
| Sequence Set | ||||
|---|---|---|---|---|
| PDB ID | Protein name | CATH Superfamily | Category | Seq. ID% to reference |
| 1OGS | Glucocerebrosidase | Glycosidases (3.20.20.80) | unfilled binding sites (reference structure) | reference str. |
| 2XWD | Glucocerebrosidase | Glycosidases (3.20.20.80) | filled binding sites | 99 |
| 2NSX | Glucocerebrosidase | Glycosidases (3.20.20.80) | filled binding sites | 100 |
| 2NT1 | Glucocerebrosidase at neutral pH | Glycosidases (3.20.20.80) | Sequence identity >60% | 100 |
| 2F7K | Pyridoxal kinase | Hydroxyethylthiazole kinase-like domain (3.40.1190.20) | unrelated <30% (identical in C) | 3 |
| 2GEP | Sulfite reductase | Adolase class I (3.20.20.70) | identical in CAT | 7 |
| 2ISB | Fumarase of FUM-1 from Archaeoglobus Fulgidus | Fumarase (3.20.130.10) | identical in CA | 10 |
| 2DJF | Human dipeptidyl peptidase I (in complex) | Cysteine proteinases (3.90.70.10) | identical in C | 7 |
| 2QGU | Phospholipid-binding protein from Ralstonia solanacearum (in complex) | Phospholipid-binding protein (1.10.10.640) | different in C | 7 |
</figtable>
Structural alignment methods results
<figtable id="struct_alis">
| PDB ID of second molecule | Pymol | SSAP | LGA | TopMatch | SAP | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| RMSD C_alpha (#atoms) | RMSD all atoms (#atoms) | RMSD (#atoms) | SSAP score | RMSD (#atoms) | LGA score | RMSD/E_r (#atoms/L) | S | S_r | Un-weighted RMSD (#atoms) | SAP score | |
| 2XWD | 0.302 (406) | 0.35 (3032) | 0.89 (492) | 95.39 | 0.75 (490) | 98.277 | 0.75 (490) | 485 | 0.71 | 0.886 (493) | 75730.101562 |
| 2NSX | 0.16 (432) | 0.196 (3383) | 0.24 (497) | 97.37 | 0.23 (497) | 100.000 | 0.23 (497) | 496 | 0.23 | 0.230 (498) | 83186.492188 |
| 2NT1 | 0.226 (454) | 0.251 (3380) | 0.68 (497) | 96.09 | 0.50 (495) | 99.287 | 0.50 (495) | 493 | 0.49 | 0.682 (498) | 76803.125000 |
| 2F7K | 18.823 (165) | 18.947 (1049) | 10.85 (177) | 48.29 | 2.92 (93) | 17.684 | 4.22 (91) | 65 | 4.02 | 16.736 (324) | 1945.130493 |
| 2GEP | 21.721 (159) | 21.9 (1006) | 4.85 (231) | 61.57 | 3.10 (60) | 7.709 | 3.07 (55) | 46 | 2.95 | 24.904 (453) | 1557.686279 |
| 2ISB | 12.845 (36) | 14.495 (245) | 14.52 (140) | 43.29 | 3.23 (55) | 16.689 | 3.09 (64) | 53 | 3.01 | 22.236 (164) | 697.835266 |
| 2DJF | 16.382 (75) | 16.896 (472) | 9.87 (90) | 41.42 | 2.98 (49) | 25.219 | 2.21 (58) | 53 | 2.11 | 11.958 (102) | 665.702637 |
| 2QGU | 22.773 (93) | 22.457 (592) | 21.03 (156) | 37.57 | 3.35 (34) | 10.973 | 2.55 (43) | 38 | 2.47 | 17.576 (176) | 559.580811 |
</figtable>
Pymol Visualisation
With Pymol we aligned the reference structure to a structure of our sequence set in <xr id="data_set"/> in two different ways. First, we aligned all atoms of both structures. Second, we focused on the alignment of the C_alpha atoms. In all cases the alignments of a structure pair only differ slightly in their composition. Moreover, in all cases the alignment of the C_alpha atoms has the lower RMSD (see <xr id="struct_alis"/>). The alignments based on C_alpha atoms, are shown in the gallery below.
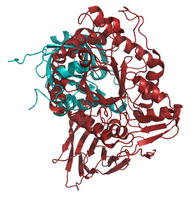
The alignment of 1OGS with 1NSX has the lowest RMSD of 0.16. Most of the C_alpha atoms are aligned. A few deviations between the structures can be observed (gallery below), however, the differences occur only in loops. The same applies for the structure 2NT1. Both structures have a 100% sequence identity to the reference structure (<xr id="data_set"/>). 2XWD, which has a sequence identity to 1OGS_A of 99%, shows indeed a very similar structure to 1OGS, differs not only in loops but also in the secondary structure. An alpha helix as well as a loop, which deviate extremely from the reference structure, can be seen marked yellow in the image.
For all other alignments, the aligned structures have nothing structural in common. Even structures that share same CATH levels are miss-aligned in their secondary structures. For structures of the first CATH level (3 Alpha Beta) at least one aligned helix or beta sheet was expected.
The main reason for that may be the very low sequence identities of these structures to Glucocerebrosidase.
- Pymol-Alignment based on C_alpha atoms: 1OGS_A (red) to structures (blue) of the set listed in Table 1
TODO(Maria): binding sites
TopMatch
TopMatch aligns only the C_alpha atoms of the structures. For the first three structures, which have a high sequence identity to our protein 1OGS_A, the length of the alignment (L) is high and the RMSD (E_r) is low (below 1). For the remaining five sequences, which have a much lower sequence identity to our protein, the RMSD becomes higher and the number of superposed residues drastically lower.
In <xr id="struct_alis"/> also the S and S_r scores are listed. The similarity score S depends on the error between each of the aligned residues and a scaling factor (sigma). The lower the error for all aligned residues, the higher S. From S, a normalized similarity per residue is calculated, dividing S by L. From s the distance error S_r is calculating (using the sigma). S and S_r are comparable to - but usually lower than - L and E_r, respectively.
SAP
The results for SAP in <xr id="struct_alis"/> show that for the sequences with a high similarity to the target, the number of superposed residues is high and the RMSD is low, similarly to TopMatch. For the sequences with a lower sequence similarity to the target, the RMSD becomes much higher (higher than on TopMatch) and the number of superposed residues lower (but remains higher than in TopMatch). This may be explained, that SAP uses the whole structure for the superposition. SAP score is very high and correlates negatively with the RMSD (the higher the score, the lower the RMSD).
Ask somebody: meaning and range of the score? Which residues are aligned?
Evaluation of structural alignments and sequence alignments
In this task, we applied the tool hhmakemodel.pl from the HHblits package to produce very crude models out of alignments by simply copying the C_alpha coordinates of the aligned residues. In this way we generated models of our protein, P04062, based on selected PDB structures found in the task 02 with HHblits. Then we evaluated how good the hhmakemodel models are aligning them to our reference structure 1OGS_A.
Results
We selected seven PDB hits for modelling with hhmakemodel. This hits were found using an HHblits search (2 iterations against uniprot20 followed by one iteration against pdb_full) with the query sequence P04062. In the first part of <xr id="hhblits_lga_scores"/> HHblits scores of sequence alignments between each hit and the query are presented: probability, E-value, score, alignment length and sequence identity. Using the PDB structures of those hits, we built structural models of the query using hhmakemodel and compared the models to the reference structure 1OGS_A using LGA. In the second part of the table the following LGA scores of structural alignments between each model and 1OGS_A are shown: number of superimposed residues, RMSD, seq_id, LGA_S and LGA_Q.
<figtable id="hhblits_lga_scores">
| HHblits sequence alignments | LGA structural alignments | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| PDB_ID | Probability | E-value | Score | Aligned_cols | Identities(%) | Superimposed residues (N) | Seq_Id(%) | RMSD | LGA_S | LGA_Q |
| 2v3f_A | 100.00 | 2.4e-132 | 1078.26 | 497 | 100 | 492 | 98.98 | 0.77 | 97.938 | 56.371 |
| 2nt0_A | 100.00 | 4.4e-132 | 1074.98 | 496 | 100 | 496 | 99.80 | 0.19 | 99.783 | 173.564 |
| 2wnw_A | 100.00 | 3.7e-107 | 870.15 | 439 | 29 | 430 | 93.49 | 1.58 | 80.275 | 25.550 |
| 3kl0_A | 100.00 | 1.1e-77 | 633.96 | 356 | 19 | 340 | 84.12 | 1.96 | 53.971 | 16.523 |
| 3s2c_A | 98.79 | 7.7e-13 | 139.02 | 246 | 14 | 221 | 87.78 | 2.49 | 29.746 | 8.524 |
| 1fob_A | 97.76 | 1.1e-08 | 101.53 | 197 | 16 | 168 | 85.71 | 2.49 | 21.221 | 6.487 |
| 1ur1_A | 95.65 | 9.8e-05 | 73.33 | 175 | 11 | 140 | 70.71 | 2.75 | 16.681 | 4.906 |
</figtable>
To see whether there is any correlation between model similarity to the reference structure and any of the alignment scores, we calculated Pearson's correlation coefficient between all pairs of the HHblits and LGA scores, which are presented in <xr id="pearsons_cc"/>.
<figtable id="pearsons_cc">
| HHblits | ||||||
|---|---|---|---|---|---|---|
| Aligned_cols | Identities(%) | Probability | E-value | Score | ||
| LGA | Superimposed residues (N) | 0.9999 | 0.8310 | 0.8530 | -0.5429 | 0.9929 |
| Seq_Id(%) | 0.8642 | 0.7958 | 0.8516 | -0.7873 | 0.8218 | |
| RMSD | -0.9372 | -0.9441 | -0.7183 | 0.4593 | -0.9392 | |
| LGA_S | 0.9690 | 0.8389 | 0.8458 | -0.5711 | 0.9383 | |
| LGA_Q | 0.6934 | 0.8304 | 0.4583 | -0.2668 | 0.6978 | |
</figtable>
The number of superposed residues of LGA alignment is strongly positive correlated to the HHblits alignment length and sequence identity, probability and score. Same correlations hold for the LGA alignment sequence identity and LGA_S score, whereas the RMDS shows strong negative correlations to the same HHblits scores. The LGA_Q score is positive correlated with these HHblits scores as well, but significantly strong only with the HHblits sequence identity. The HHblits E-value has the opposite correlation tendencies to the LGA scores than the other HHblits scores. It is strongly negative correlated to the LGA sequence identity, medium negative correlated to the number of superposed residues of LGA alignment and LGA_S, medium positive correlated to the RMSD and slightly negatively correlated to LGA_Q. This means, that HHblits hits with a high alignment length and sequence identity, probability and score lead to good quality of hhmakemodel models (with high number of superposable residues, sequence identity and scores and low RMSD to the reference structure). Low E-value of HHblits hits also contributes to the model quality.