Difference between revisions of "Homology Modelling GLA"
(→Comparison to Experimental Structure) |
(→Comparison to Experimental Structure) |
||
| Line 121: | Line 121: | ||
===Numeric Evaluation=== |
===Numeric Evaluation=== |
||
===Comparison to Experimental Structure=== |
===Comparison to Experimental Structure=== |
||
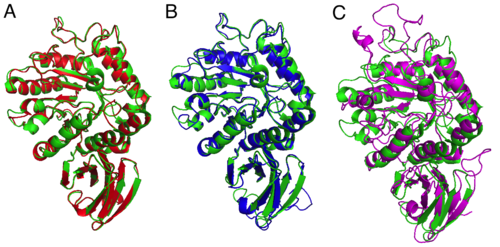
| + | [[File:Fabry_disease_homology_modelling_modeller_combined.png|thumb|500px|center|Representation of the resulting models of MODELLER and the reference PDB structure [http://www.rcsb.org/pdb/explore/explore.do?pdbId=1r47 1R47]. The models are in superposition to the reference structure (green) and are shown in cartoon representation. (A) The model is based on the PDB structure 3HG3 (red). (B) The model is based on the PDB structure 1KTB. (C) The model is based on the PDB structure 3CC1.]] |
||
| − | {| class="centered" |
||
| − | |[[File:Fabry_disease_homology_modelling_modeller_combined.png|thumb|Graphical output of TMHMM for the protein GLA.]] |
||
| − | |} |
||
==iTasser== |
==iTasser== |
||
Revision as of 14:14, 14 June 2011
by Benjamin Drexler and Fabian Grandke
Calculation of Models
Available Homologous Structures
The HHpred search from Task1 output the following sequences(Top 10):
| PDB-ID | Name | Probability | E-value | P-value | Identity |
|---|---|---|---|---|---|
| > 60% sequence identity | |||||
| 3hg3_A | Alpha-galactosidase A | 1.0 | 0 | 0 | 97% |
| > 40% sequence identity | |||||
| 1ktb_A | Alpha-N-acetylgalactosaminidase | 1.0 | 0 | 0 | 53% |
| < 40% sequence identity | |||||
| 1uas_A | Alpha-galactosidase | 1.0 | 0 | 0 | 39% |
| 3lrk_A | Alpha-galactosidase 1 | 1.0 | 0 | 0 | 32% |
| 3a5v_A | Alpha-galactosidase | 1.0 | 0 | 0 | 35% |
| 1szn_A | Alpha-galactosidase | 1.0 | 0 | 0 | 34% |
| 3a21_A | Putative secreted alpha-galactosidase | 1.0 | 0 | 0 | 34% |
| 3cc1_A | BH1870 protein | 1.0 | 0 | 0 | 26% |
| 3a24_A | Alpha-galactosidase | 1.0 | 0 | 0 | 14% |
| 1zy9_A | Alpha-galactosidase | 1.0 | 2.2E-37 | 8.8E-42 | 14% |
MODELLER
We used the MODELLER as described in the tutorial Using Modeller for TASK 4.
| PDB-ID | Unsupervised | Supervised | Identity | Comment |
|---|---|---|---|---|
| 3LX9_A | 99% | |||
| 3GXP_A | 99% | |||
| 3H53_A | 99% | |||
| 3HG3_A | 97% | |||
| 3IGU_A | 54% | |||
| 1KTB_A | 53% | |||
| 1UAS_A | 39% | |||
| 3LRK_A | 34% | Was removed due to little sequence identity. Caused huge gaps in alignment. | ||
| 3CC1_A | 28% | Was removed due to little sequence identity. Caused huge gaps in alignment. |
iTasser
Figure 1 shows, that iTasser takes an amino acid sequence as input and tries to retrieve template proteins from PDB. In the next step fragments from the the templates are reassembled to a complete model. In the last step, the model is reassembled by taking energy calculations into account. Additionally biological function prediction is done, but that was not of interest of this task.<ref name=itasser1>http://zhanglab.ccmb.med.umich.edu/I-TASSER/about.html</ref>

We used the iTasser-server in two different ways:
- Standard parameters: the protein sequence is given as input and the program searches PDB for templates. The found proteins are used to create a template to predict the structure.
- PDB-ID as input: together with the amino acid sequence a template PDB-ID is given as input. The program takes all available information into account and uses them to calculate the structure.
As the iTasser server has very low capacities and only one job commitment at the same time is possible, the results of the second way are not yet present. The standalone version is no option, because it has a size of about 10GB and it does not work properly.
SWISS-MODEL
We used the swissmodel server with two different options:
- Automated Mode: A template sequence is given as input. As no further information are given, the model is directly created from the amino acid sequence. This method should only be used, if the sequence identity between target and template is greater than 50%.
- Aligned Mode: A pairwise alignment of template and target sequence is given as input. We created our alignments using online ClustalW2 from EBI.
Following sequences have been selected:
| 3hg3_A | 1ktb_A | 3cc1_A | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Automated Mode | Aligned Mode | Automated Mode | Aligned Mode | Automated Mode | Aligned Mode | |||||||||
| Identity | Z-score | Model | Z-score | Model | Identity | Z-score | Model | Z-score | Model | Identity | Z-score | Model | Z-score | Model |
| 97% | 0 | -0.415 | 53% | 0 | -2.261 | 26% | 0 | Error¹ | NA | |||||
¹The sequences are to different to create a useful model from the alignment. In the automated mode the template itself has been used as model, what is useless, because the sequences have only 26% identity.
Aborting: too many unfruitful attempts to rebuild a loop.
This is likely to indicate a misalignment in this region
Use Swiss-PdbViewer to adjust your alignment in this
region and resubmit an optimise mode modelling request.
Evaluation of Models
MODELLER
Numeric Evaluation
Comparison to Experimental Structure
iTasser
Numeric Evaluation
Comparison to Experimental Structure
SWISS-MODEL
Numeric Evaluation
Comparison to Experimental Structure
References
<references />