Difference between revisions of "Sequence Alignments Hemochromatosis"
Bernhoferm (talk | contribs) (→Blast) |
Bernhoferm (talk | contribs) (→Blast) |
||
| Line 42: | Line 42: | ||
</figtable> |
</figtable> |
||
| − | The first BLAST search against the Big80 database reached the hit limit of 250 sequences with an |
+ | The first BLAST search against the Big80 database reached the hit limit of 250 sequences with an e-Value of e-30 for the worst hit. So we did it again with a new limit of 1500 reported hits. These hits were then filtered for unique IDs and an e-Value cutoff of 2e-3. After the filtering 1159 hits were left. |
| − | The distributions for the |
+ | The distributions for the e-Values and identities of these hits are shown in <xr id="blastdist"/>. Most of the e-Values are between 1e-50 and 2e-3 (cutoff). Only few hits have a better e-Value. The identities are piled between 20% and 40% with two peaks at around 27% and 34% respectively. |
=== PSI-Blast === |
=== PSI-Blast === |
||
Revision as of 10:03, 7 May 2012
Henry Frankenstein: Look! It's moving. It's alive. It's alive... It's alive, it's moving, it's alive, it's alive, it's alive, it's alive, IT'S ALIVE!
Victor Moritz: Henry - In the name of God!
Henry Frankenstein: Oh, in the name of God! Now I know what it feels like to be God!
Contents
Task Description
Protocol
Sorry for the inconvenience (not beeing able to read something), we're rerunning some data...
Reference Sequence
Sequence from Uniprot: Q30201
>sp|Q30201|HFE_HUMAN Hereditary hemochromatosis protein OS=Homo sapiens GN=HFE PE=1 SV=1 MGPRARPALLLLMLLQTAVLQGRLLRSHSLHYLFMGASEQDLGLSLFEALGYVDDQLFVF YDHESRRVEPRTPWVSSRISSQMWLQLSQSLKGWDHMFTVDFWTIMENHNHSKESHTLQV ILGCEMQEDNSTEGYWKYGYDGQDHLEFCPDTLDWRAAEPRAWPTKLEWERHKIRARQNR AYLERDCPAQLQQLLELGRGVLDQQVPPLVKVTHHVTSSVTTLRCRALNYYPQNITMKWL KDKQPMDAKEFEPKDVLPNGDGTYQGWITLAVPPGEEQRYTCQVEHPGLDQPLIVIWEPS PSGTLVIGVISGIAVFVVILFIGILFIILRKRQGSRGAMGHYVLAERE
Sequence Searches
Blast
<figtable id="blastdist">
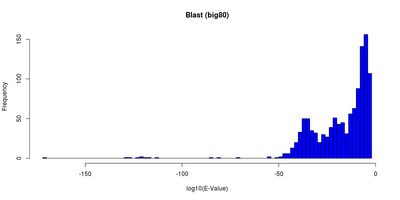
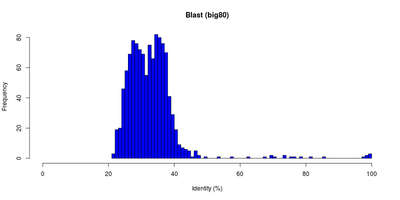
| Table 1: E-Value and identity distributions of the Blast search within Big80. |
</figtable>
The first BLAST search against the Big80 database reached the hit limit of 250 sequences with an e-Value of e-30 for the worst hit. So we did it again with a new limit of 1500 reported hits. These hits were then filtered for unique IDs and an e-Value cutoff of 2e-3. After the filtering 1159 hits were left.
The distributions for the e-Values and identities of these hits are shown in <xr id="blastdist"/>. Most of the e-Values are between 1e-50 and 2e-3 (cutoff). Only few hits have a better e-Value. The identities are piled between 20% and 40% with two peaks at around 27% and 34% respectively.
PSI-Blast
<figtable id="psiblastdist">
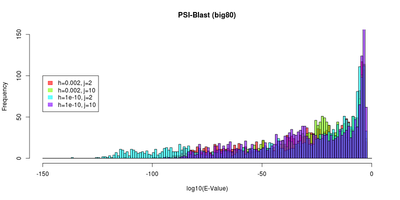
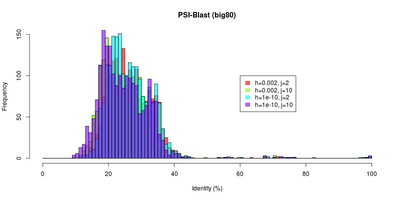
| E-Value and identity distributions of the PSI-BLast search within Big80. |
</figtable>
HHblits
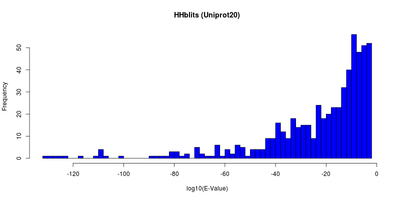
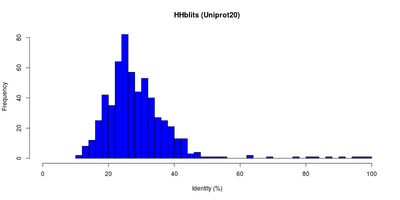
<figtable id="hhblitsdist">
| E-Value and identity distributions of the HHblits search within Uniprot20. |
</figtable>
Comparison
<figtable id="psiblastoverlap">
| Nothing to see here.... |
</figtable>
<figtable id="alloverlap">
| Nothing to see here.... |
</figtable>