ASPA Homology Modelling
Contents
Homology Modelling
Selected Structures
We used HHSearch and NCBI related structure search to search for usable structures. We applied a threshold of alignments of at least 150bp length to weed out the myriad of hits with good sequence similarity but extremely short alignments.
In the end, we got the following list of structures:
| PDB ID | Sequence Identity | Alignment Length | Description |
|---|---|---|---|
| > 60% sequence identity | |||
| 2GU2_A | 87% | 307 | Aspartoacyclase, Rattus norvegicus |
| > 40% sequence identity | |||
| 3NH4_A | 43% | 309 | Aspartoacyclase, Mus musculus |
| > 0% sequence identity | |||
| 2QJ8_A | 26% | 178 | MLR6093 Hydrolase, Mesorhizobium loti |
| 1YW4_A | 22% | 180 | Succinylglutamate desuccinylase, E. coli |
| 3CDX_A | 18% | 180 | Succinylglutamate desuccinylase, Rhodobacter sphaeroides |
The list is rather thin in the upper percentage ranges, but no other structures could be found. In the bottom region, another Succinylglutamate desuccinylase was omitted, since it had an even worse alignment length and identity percentage and a cluster of closely related proteins of very low sequence identity was thought to slant the prediction in undesirable directions.
Single Template Modelling
Modeller
We chose the structures 2GU2, 3NH4 and 2QJ8 as our templates, since they were either without alternative or best in their group. With standard parameters, we got the following models; in one case, modeller tried to align our target onto the while homodimer sequence of the template, which resultet in nonsens. In accord with the instructions, we didn't do any optimization in this step; we did enforce use of one specified chain later.
We found that in the case of 2GU2, no editing was necessary. The non-SSE-aware alignment aligned the target nicely with the B chain (which is pretty much identical to the A chain and therefore OK); there were no gaps in functional regions or any other issues, so we left it untouched.
The same was true for 3NH4; what gaps we found were either very sensible or without import. Consequently, we did not modify this alignment either.
In the case of 2QJ8, not entirely unexpected problems arose; here, modeller's alignment algorithm had chopped the target sequence into many small segments separated by long stretches of gaps in a futile attempt to produce a global alignment over the whole homodimer sequence. We solved this by building a new PDB file with one chain removed and re-constructing the model. The result was a marked improvement on the previous attempt:
SwissModeller
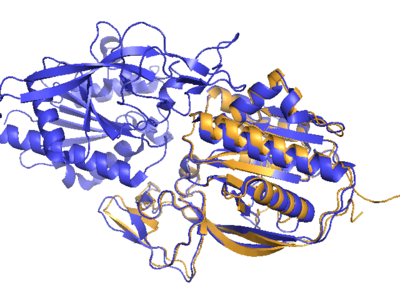
We also submitted our target and the three selected templates to SwissModeller. Superposited images of the resulting structures versus the real experimental one are shown on the right of this page.
iTasser
We could not complete this step, because iTasser stated that it had a very very long back queue and would process our jobs once all pending jobs were done; that was all we ever received from it. We will append the structures to this document if iTasser should ever happen to finish our jobs, however unlikely a prospect that may be after all the waiting we've already done.
Multiple Template Modelling
We also tried using all three templates from the third identity segment to construct one model. We let Modeller construct an MSA by itself and tried to edit it, quickly giving up since we could not effect any visible improvements and, by contrast, even produced models that looked worse on visual inspection in PyMol.
This step returned the following structure:
Evaluation
For superposition images the structure documented on our disease Wiki page, 2O53, was used. The pictures are shown on the right.
| Model | RMSD | TM Score |
|---|---|---|
| Modeller edited 2QJ8 | 2.304 | 0.5024 |
| Modeller Multi-template | 0.427 | 0.9801 |
| Modeller standard 2GU2 | 0.412 | 0.9858 |
| Modeller standard 2QJ8 | 2.467 | 0.2706 |
| Modeller standard 3NH4 | 0.776 | 0.9662 |
| SwissModel 2GU2 | 0.413 | 0.9787 |
| SwissModel 2QJ8 | 1.746 | 0.5494 |
| SwlssModel 3NH4 | 0.769 | 0.9651 |
Discussion
The best method is modeller with the 2GU2 template; since the similarity between 2GU2 and the target is extremely high, it is hardly surprising that one of these models happens to be the best one generated; what does come as a surprise is that Modeller outperforms SwissModel for this structure, albeit only by a very small margin. Also very good performance was reached by using multiple templates with Modeller; here, the result is almost as good as that which used the 2GU2 template. This is interesting, as the templates used were of very bad quality, with the one most similar to the target, 2QJ8, leading to extremely bad results when used as single template. Here Modeller outperforms SwissProt again, although this time in screwing up badly.
Modeller appears to lean more towards extremes; its extreme cases are both better and worse than thouse of SwissModel. Sequence similarity played a big role, with 2GU2 being always the best one by a comfortable margin, 3NH4 second best and 2QJ8 pretty much catastrophic. That this can be compensated by combining information from three bad structures is quite fascinating.