Normal mode analysis (Phenylketonuria)
From Bioinformatikpedia
Page still under construction!!!
Contents
Summary
...
Normal Mode Analysis
Lab journal
For the analysis, we used the pdb structure 1J8U from the task before (Task 9 - Structure-based mutation analysis).
WEBnm@
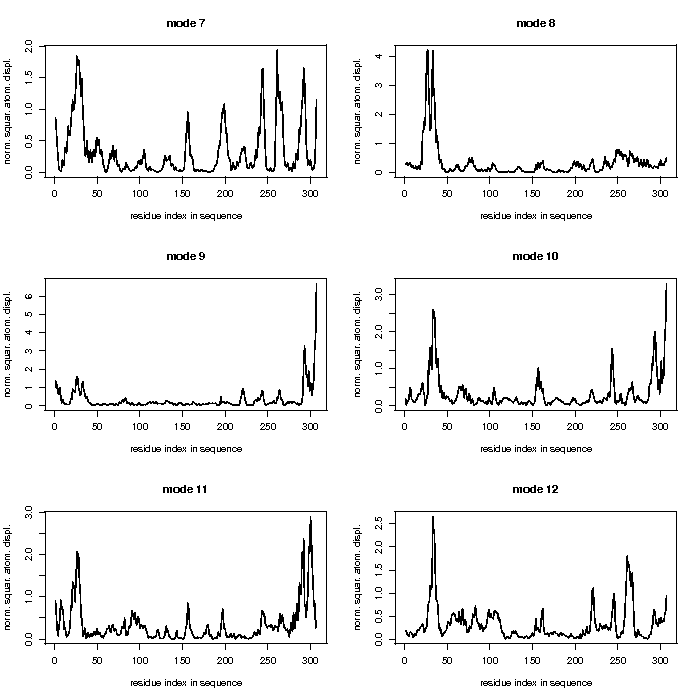
Mode 7
...
Mode 8
...
Mode 9
...
Mode 10
...
Mode 11
...
Mode 12
...
elNémo
ElNémo computes models for the given protein 1J8U. Thereby it calculates 100 lowest-frequency modes and provides a lot of information about the modes with different parameters and visualizations:
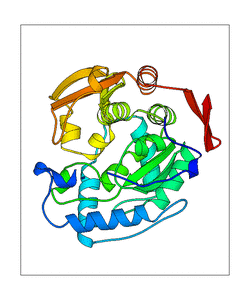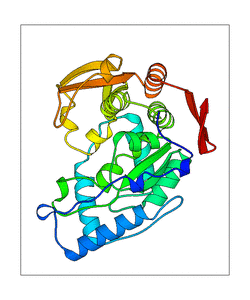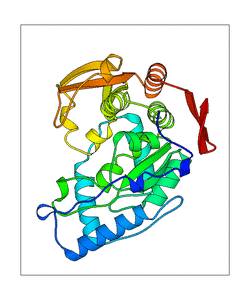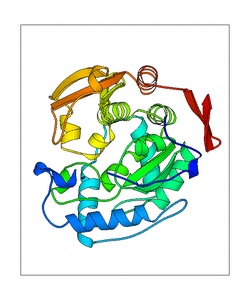
- PDB-files with the conformations of the modes
- animations of the modes of three different sites
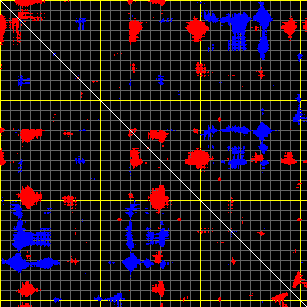
- CA-vari (calculates distance fluctuations between all C-alpha atoms)
- R2 residue mean square displacement of all C-alpha atoms
- Frequency and collectivity of the modes
<figtable id="frecol">
| First five modes | |||
|---|---|---|---|
| Mode | Frequency | Collectivity | |
| mode 7 | 1.00 | 0.5475 | |
| mode 8 | 1.02 | 0.4062 | |
| mode 9 | 1.36 | 0.2941 | |
| mode 10 | 1.43 | 0.0948 | |
| mode 11 | 1.60 | 0.3466 | |
</figtable>
- B-factor analysis for the correlation between observed and normal-mode-derived atomi displacement parameters:
- Correlation = 0.534 for 307 C-alpha atoms
- RMSD: only if two structures are used
The analyses of the five modes with lowest normal modes are shown below.
Mode 7
Mode 8
Mode 9
Mode 10
Mode 11
VMD
...
References
<references/>