Task 6 (MSUD)
Contents
Results
H-Ras
Freecontact
Corrected norm contact (CN) scores are gathered from pairs of residues which have larger than 5 residues in between. Distribution of CN scores shows a normal-like distribution where most residue pairs have low contacts and few residues have very significant contacts between them. The CN scores from prediction over H-Ras protein range from -0.65 to 5.99.
After fitting the CN scores to a normal distributes (μ = 3.7×10-4, σ=0.494), we choose the 98-th percentile (≈1.0) as cutoff for significant contacts. With the R script by Laura, 63 high-scoring contacts were found. In comparison to the protein structure 121P 54 contact pairs are categorized as true positives. Following are the top-10 contact pairs with highest CN scores:
| 1. residue # | 1. aa | 2. residue # | 2. aa | MI score | CN score | true/false positive |
|---|---|---|---|---|---|---|
| 11 | A | 92 | D | 0.32 | 3.40 | TP |
| 81 | V | 116 | N | 0.24 | 3.00 | TP |
| 87 | T | 129 | Q | 0.22 | 2.68 | FP |
| 19 | L | 81 | V | 0.15 | 2.55 | TP |
| 82 | F | 141 | Y | 0.25 | 2.53 | TP |
| 84 | I | 115 | G | 0.16 | 2.52 | TP |
| 82 | F | 115 | G | 0.14 | 2.42 | TP |
| 10 | G | 16 | K | 0.39 | 2.24 | TP |
| 130 | A | 141 | Y | 0.39 | 2.23 | TP |
| 123 | R | 143 | E | 0.27 | 2.20 | TP |
After computing and sorting high scoring residues, we have found 4 different hot spots, which seems to be responsible to the stability of structure of H-Ras. They are the Phe 82, Val 81, Ala 11 and Tyr 40. The residues contribute to the structural stability are marked as fallowing:
- Hotspots in structure of H-Ras protein
EVcoupling
| 1. residue # | 1. aa | 2. residue # | 2. aa | MI score | CN score | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 4 | Y | 5 | K | 0.505568 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 6 | L | 0.22551 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 7 | V | 0.327402 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 8 | V | 0.198472 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 9 | V | 0.2039 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 10 | G | 0.117158 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 11 | A | 0.374551 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 12 | G | 0.310161 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 13 | G | 0.252042 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | Y | 14 | V | 0.192398 | }
EVfoldBCKDHAFreecontactFrom the output of freecontact, only those residue pairs were extracted, that are separated by at least 5 residues. Pairs that are near in sequence are natually coupled, but this isn't interesting because it doesn't help to predict the 3D structure, where residues intereact, which are far away from each other in the 1D sequence. The CN (corrected norm contact) score values of these extracted pairs range between -0.87 and 5.28. The following diagram shows the distribution of the scores: Most scores are between -1 and 1. So those pairs, which have a score greater than 1 are considered high scoring and used for further analyses. High scoring pairs were regarded as true positive (TP), if their distance (between any pair of atoms) in the reference structure is below 5 Å. There are 94 TPs among 194 high scoring pairs, and this has a correlation to CN score of 0.31 (for a pair with higher CN score it is more likely that it is TP). This table shows the ten highest scoring pairs:
EVcouplingsThe following table shows the ten residue pairs with highest DI (direct information) scores:
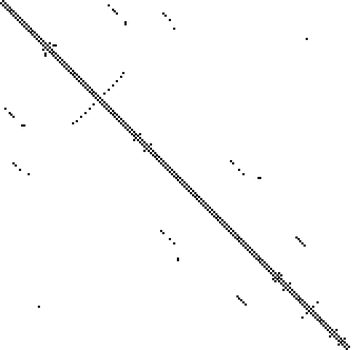
EVfoldFor every number of contraints, EVfold computes 5 different structure models. The RMSD values for these models compared to the reference structure 1U5B are summarized in the following boxplot (RMSD values taken from EVfold output): For 195 contraints (corresponding to 65 % of alignment length) the models agree more with the known structure, than if only 120 (40 %) contraints are used. To use 300 (100 %) constraints doesn't help for structure prediction compared to 65 %, it seems to work even worse for structure prediction (although there is a large range of RMSDs between the different models calculated withh 300 contraints). In general, the RMSD values are very high, indicating that the models are very different to the real structure. The contact maps show the predicted contacts compared to the real contacts in 1U5B (the calculation of EVfold doesn't cover the whole sequence): For the lowest number of constraints, most are true positives, but some regions with contacts in the known structure are only covered, if more constraints are used. But in the second and third plot, it is visible that when using more constraints, there are more false positives (predicted contacts where in the real structure there is no contact). So using 60-70 % contraints may be a good compromise between recall and precision. Discussion
|