MD Mutation485
Contents
- 1 check the trajectory
- 2 Visualize in pymol
- 3 create a movie
- 4 energy calculations for pressure, temperature, potential and total energy
- 5 minimum distance between periodic boundary cells
- 6 RMSF for protein and C-alpha
- 7 Radius of gyration
- 8 solvent accesible surface area
- 9 hydrogen-bonds
- 10 Ramachandran plot
- 11 RMSD matrix
- 12 cluster analysis
check the trajectory
We checked the trajectory with following command:
gmxcheck -f mut436_md.xtc
With the command we got following results:
Reading frame 0 time 0.000 # Atoms 96545 Precision 0.001 (nm) Last frame 2000 time 10000.000
Furthermore, we got some detailed results about the different items during the simulation.
| Item | #frames | Timestep (ps) |
| Step | 2001 | 5 |
| Time | 2001 | 5 |
| Lambda | 0 | - |
| Coords | 2001 | 5 |
| Velocities | 0 | - |
| Forces | 0 | - |
| Box | 2001 | 5 |
The simulation finished on node 0 Thu Sep 15 19:12:47 2011.
| Time | ||
| Node (s) | Real (s) | % |
| 22336.000 | 22336.000 | 100% |
| 6h12:00 | ||
The complete simulation needs 6 hours and 12 minutes to finishing.
| Performance | |||
| Mnbf/s | GFlops | ns/day | hour/ns |
| 1277.617 | 93.808 | 38.682 | 0.620 |
As you can see in the table above, it takes about 2/3 hour to simulate 1 ns of the system. So therefore, it would be possible to simulate about __ns in one complete day calculation time.
Visualize in pymol
First of all, we visualized the simulation with with ngmx, because it draws bonds based on the topology file. ngmx gave the user the possibility to choose different parameters. Therefore, we decided to visualize the system with following parameters:
| Group 1 | Group 2 |
| System | Water |
| Protein | Ion |
| Backbone | NA |
| MainChain+H | CL |
| SideChain |
Figure 1 shows the visualization with ngmx:
create a movie
Next, we want to visualize the protein with pymol. Therefore, we extracted 1000 frames from the trajectory, leaving out the water and jump over the boundaries to make continuse trajectories. Therefore, we used following command:
trjconv -s fole.tpr -f file.xtc -o output_file.pdb -pbc nojump -dt 10
The program asks for the a group as output. We only want to see the protein, therefore we decided to use group 1.
Todo: film und filtered
energy calculations for pressure, temperature, potential and total energy
Pressure
| Average (in bar) | 0.998385 |
| Error Estimation | 0.0058 |
| RMSD | 71.0317 |
| Tot-Drift | -0.0436306 |
The plot with the pressure distribution of the system can be seen here:
As you can see in Figure 5, the pressure fluctuates in the system around 0 and does mostly not exceed 100. Contrary,there are some outlier which reach values of almost 250 and -250 bar. In this cases we are not sure, if a protein works under such a high pressure.
Temperature
| Average (in K) | 297.936 |
| Error Estimation | 0.0045 |
| RMSD | 0.940566 |
| Tot-Drift | 0.00654126 |
The plot with the temperature distribution of the system can be seen here:
Figure 2 displays the distribution of the temperature in the current MD system. In this case the temperature fluctuates around 298K. The maximal occuring amplitude between the average and the outlier temperature is only about 3 K which is not much. But we have to keep in mind, that only some degree difference can affect huge functional loss of a protein. 298 K corresponds to 25°C, which is relatively low for human protein activity, because the highest activity is normally reached at body temperature which is about 37°C.
Potential
| Average (in kJ/mol) | -1.28176e+06 |
| Error Estimation | 85 |
| RMSD | 1068.67 |
| Tot-Drift | -536.314 |
The plot with the potential energy distribution of the system can be seen here:
Figure 3 displays the potential engery of the system which is between -1.285e+06 and -1.275e+06. This is a relativly low energy. Therefore, this probably means that the protein is stable. Furthermore, we can suggest, that the protein with such a low energy is able to function and is stable. Therefore, our simulation is probably true. Otherwise, if the energy of the simulated system would be too high, we can not trust the results, because the protein is too instable to work.
Total energy
| Average (in kJ/mol) | -1.05203e+06 |
| Error Estimation | 83 |
| RMSD | 1308.04 |
| Tot-Drift | -531.275 |
The plot with the total energy distribution of the system can be seen here:
Looking at Figure 4, we can see that the total energy deviates a little from the potential energy which is minimal lower. In this case, the total energy is between -1.055e+06 and -1.045e+06. These values lie already in a range, where we can suggest that the protein energy is sufficient low so that this one can work.
minimum distance between periodic boundary cells
Next we try to calculate the minimum distance between periodic boundary cells. As before, the program asks for one group to use for the calculation and we decided to use only the protein, because the calculation needs a lot of time and the whole system is significant bigger than only the protein. So therefore, we used group 1.
Here you can see the result of this analysis:
- Hier noch ändern: ***************************
Figure 6 and 7 display.... , there is a huge difference between the different time steps and distances. The highest distance is up to 4 nm, whereas the smallest distance is only about 1nm. Therefore, we can see that the protein is very flexible over the time.
RMSF for protein and C-alpha
Protein
First of all, we calculate the RMSF for the whole protein.
The analysis produce two different pdb files, one file with the average structure of the protein and one file with high B-Factor values, which means that the high flexbile regions of the protein are not in accordance with the original PDB file.
To compare the structure we align them with Pymol with the original structure.
| original & average | original & B-Factors | average & B-Factors |
| Perspective one | ||
| Perspective two | ||
| RMSD | ||
| 1.519 | 0.349 | 1.727 |
- Hier noch ändern: ***************************
The structure with the high B-factors is the most similar structure (Figure 8 and Figure 11) compared with the original structure from PDB (Figure 7 and Figure 10). The average structure is not that similar (Figure 10 and Figure 12). But we know, that the regions with high B-Factors are very flexible, and therefore in the structure downloaded from the PDB, the protein is in another state, because of its flexible regions. Therefore, because of the low RMSD between the high B-factors structure and the original structure we can see, that the simulation predicts the structure quite good.
Furthermore, we got a plot of the RMSF values of the protein, which can be seen in Figure 13:
- Hier noch ändern: ***************************
There are two regions with very high B-factor values. One region at position 150 (Figure 14), and the other region at position 440 (Figure 15). If we compare the picture of the original and the average structure, we can see that most of the regions build a very good alignment, whereas some regions vary in their position. Therefore, we want to compare, if these regions are the regions with very high B-factor values.
Furthermore, we visualized the B-factors with the Pymol selection B-factor method. We calculated the B-factors for the blue protein (Figure 16 and Figure 17). If you see red, this part of the protein is very flexible. The brighter the color, the higher is the flexibility of this residue.
- Hier noch ändern: ***************************
In the second picture, you can see, that the color is dark blue. Therefore a peak lower than 0.3 do not mean that there is high flexibility. Therefore, our protein has only one very flexible region and this is around residue 140.
As you can see in the pictures above, especially in the first picture, which is the part with the highest peak in the plot, the structures have a very different position and the alignment in this part of the protein is very bad, although the rest of the alignment is quite good. This also explains the relatively high RMSD values, because of the different positions of the flexible parts of the protein.
C-alpha
Now we repeat the analysis done for the protein for the C-alpha atoms of the protein. Therefore, we followed the same steps as in the section above.
To compare the structure we align them with pymol with the original structure.
| original & average | original & B-Factors | average & B-Factors |
| Perspective one | ||
| Perspective two | ||
| RMSD | ||
| 1.258 | 0.283 | 1.297 |
- Hier noch ändern **********************
As in the section above, the RMSD between the structure with high B-factor values and the original structure is the most similar (Figure 19 and Figure 22). This was expected, because we used twice the same model, but in this case we neglecte the residues of the atoms. But the backbone of the protein remains the same. The other two models (Figure 18, Figure 20, Figure 21 and Figure 23) have nearly the same RMSD value and therefore there are equally.
Furthermore, we got a plot of the RMSF values of the protein, which can be seen on Figure 24:
- Hier noch ändern **********************
In this case, there is only one high peak at position 150. By observing the whole protein, it was possible to see, that the position of the beta sheet differs extremely between the two models. The other peak at position 440 could not be found in the plot. By a look at the picture above, we can see that the backbone do not differ extremely between the two models. Therefore, in this case the position of the residues has to be very different, which is not important in our case, because we do not regard side chains.
Radius of gyration
Next, we want to analyse the Radius of gyration. Therefore we use g_gyrate and use only the protein for the calculation.
- Hier noch ändern **********************
Figure 25 shows the radius of gyration over the simulation time. The Radius of gyration is the RMS distance of its parts from its center or gyration axis. On the plot it could be seen, that the average radius is about 2.4, but there are big differences, which means, that the protein is flexible. The distance between the different parts of the protein and the center differs so the protein seems to pulsate, because there is a periodic curve which shows the loss and the gain of space the protein needs.
solvent accesible surface area
Next, we analysed the solvent accesible surfare area of the protein, which is the area of the protein which has contacts with the surronding environment, mainly water.
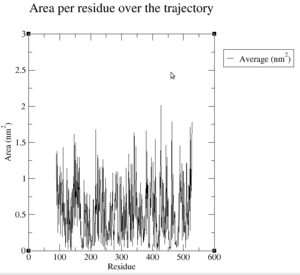
First of all, we have a look at the solvent accesibility of each residue, which can be seen on Figure 26.
Furthermore, it is also possible to look at the solvent accesibility of each atom of the complete protein, which can be seen on Figure 27.
Figure 29 shows how much of the area of the protein is accesibile to the surface during the complete simulation. As we saw before, by the gyration radius of the protein, the values differ during the simulation, which shows, that the protein is flexible.
- Hier noch ändern **********************
So we can see in the pictures there are a lot of differences between the solvent accesibility of the protein's surface, which was expected. If we have a closer look at the plot about the atomic accesibility and the residue accesibility, we can see, that both pictures agree. During the simulation there is a big differences in the accessibility of the protein. There is a area between 131 and 146 nm/2S/N during the simulation, which is a big difference. Therefore, we can see that the protein has to be really flexible, otherwise, these different values could not be explained.
hydrogen-bonds
In this case, we differ between hydrogen-bonds between the protein itselfs and bonds between the protein and the water.
As before, it is possible to see in the plot, that the protein is flexible, because the number of bonds differ extremely over the time.
On Figure 30 you can see the bonds between the protein. Here you can see that the number differs between 300 bonds and 355. Most of the time, the protein has between 320 and 330 hydrongen-bonds.
- Hier noch ändern **********************
If we have a look at the number of bonds between the protein and the water, which are visualized on Figure 31, we can see that there are a lot more bonds between protein and water than in between the protein. The number differs between 800 and 900 and there there are about 3 times more bonds between the protein and the water. Over the simulation time, the number of bonds between water and protein grows in average. But most of the time, the protein has between 840 and 860 bonds with the water.
- Hier noch ändern **********************
This is no surprise, because every residue on the surface has contact with water, whereas in the protein there are a lot of amino acids which do not have contact partners, because the other amino acids are too far away.
Ramachandran plot
Now, we want to have a closer look to the secondary structure of the protein during the simulation. Therefore, we used a Ramachandran plot to analyse the phi and psi torsion angles of the backbone to get a better understanding of the secondary structure during the simulation.
- Hier noch ändern **********************
As we can see on Figure ?, there are a lot of beta sheets, alpha helices and right-handed alpha helices. The white regions are the regions where no secondary structure can be found, which is right.
RMSD matrix
Next we analysed the RMSD values. Therefore, we used a RMSD matrix. This is useful to see if there are groups of structures over the simulation that share a common structure. These groups will have lower RMSD values withing their group and higher RMSD values compared to structure which are not in the group.
The following matrix shows the RMSD values of our structures.
- Hier noch ändern **********************
As you can see in the picture above, there is one big group which is colored in green, but it is not possible to find any very dense groups which all have a very low RMSD compared to each other. Only near on the diagonal there are some structures which are colored in cyan, but in general, most of the structures are colored in green which means a RMSD value of about 0.15 Angstrom. So we can see that there are differences of the protein structure during the simulation and therefore we can conclude that this protein is very flexible.
cluster analysis
Next, we started a cluster analysis. First of all, we found 225 different clusters.
We visualized all of these cluster structures:
Next we aligned some structures of the cluster and measured the RMSD:
| Cluster 1 | Cluster 2 | RMSD |
| cluster 1 | cluster 2 | |
| cluster 1 | cluster 5 | |
| cluster 30 | cluster 100 |
- Hier noch ändern **********************
The RMSD values of the different structures are very similar, which can be seen in the picture above. Furthermore, the RMSD values of the different structures of the clusters are very low. But if we align structures which are far away in number, the RMSD value increase, which shows us that over the simulation time the differences between the start structure and the simulated structures increase. Therefore, we can see that the different structures of the simulation are very similar, but over the time they change more and more.
To have a better insight into the distribution of the RMSD value between the different clusters, we visualize the distribution in Figure ?.
- Hier noch ändern **********************
On Figure ?, it is possible to see, that the distribution is a gaussian distribution, with the highest peak at 0.2 Angstrom. This means, that most of the structures have a RMSD about 0.2 Angstrom compared to the start structure. This value is relatively high, especially if we keep in mind, that both structure are the same structure, and only for one structure the motion over the time was simulated.