Sequence-based mutation analysis (Phenylketonuria)
Contents
Summary
...
Mutation dataset
| SNPs | |||
|---|---|---|---|
| AA - three letter | AA - one letter | Nucleotides | |
| Ala259Val | A259V | C776T | |
| Arg123Ile | R123I | G368T | |
| Gln20Leu | Q20L | A59T | |
| Gln172His | Q172H | G516T | |
| Gly103Ser | G103S | G307A | |
| Ile421Thr | I421T | T1262C | |
| Lys341Thr | K341T | A1022C | |
| Phe392Ser | F392S | T1175C | |
| Pro416Gln | P416Q | C1247A | |
| Thr266Ala | T266A | A796C | |
Analyze SNPs
The complete results of the prediction tools can be found in the Lab journal.
Looking at the substitution matrices you can see that the substitutions analyzed in this task never have the worst values.
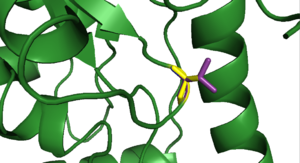
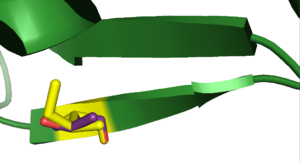
As 2pah begins at position 118 for amino acids 20 and 103 the substitutions cannot be shown with pymol. A comination of the other eight mutations can be seen in <xr id="all_mutations"/>. None of those amino acids participates in forming the binding site, so it is not disturbed by any of the substitutions that are examined here.
<figure id ="all_mutations">
</figure>
Ala-259-Val
<figure id="A259V">
</figure>
| Properties | alanine is small, non-polar, neutral and hydrophobic |
| valine is aliphatic, small, non-polar, neutral and hydrophobic | |
| Structure | LOOP |
| Substitution Matrices | middle ranged value → neutral substitution |
| PSSM | alanine seems to be highly conserved with a value of 70. Valine has a value of 1. |
| SIFT | TOLERATED with a score of 0.16 |
| PolyPhen2 | probably damaging with a score of 1.000 |
| SNAP | non-neutral, RI = 2 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
For the substitution of alanine to valine the properties do not change very much as the only difference is that valine is aliphatic. This can also be seen in the substitution matrices where such an exchange has a middle ranged value and therefore indicates a neutral substitution. Additionally SIFT predicts this mutation to be tolerable, but the score is not very high. When looking at the PSSM matrix that was created with psiblast a high conservation of alanine on position 259 can be seen, where valine only has a low tolerable value here. Furthermore the other three prediction tools all predict a non-neutral effect of this substitution where most notably PolyPhen2 has a very high score. SNAP only has a reliability index of two, but an accuracy of 70%. Looking at the structure itself (<xr id="A259V"/>) you can see that the substitution is located in a LOOP, however it is directly beneath a HELIX or the last amino acid of it, which is predicted to get lost by MutationTaster if the amino acid is changed. In this case, to decide if the substitution is neutral or not is really hard as the different features suggest both. However, we predict this mutation to be non-neutral.
Arg-123-Ile
<figure id="R123I">
</figure>
| Properties | arginine is positively charged, polar and hydrophilic |
| isoleucine is aliphatic, neutral, non-polar and hydrophobic | |
| Substitution Matrices | low value → bad substitution |
| Structure | LOOP |
| PSSM | arginine seems to be highly conserved with a value of 49. Isoleucine has a value of 7. |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.00 |
| PolyPhen2 | possibly damaging with a score of 0.807 / 0.582 |
| SNAP | neutral, RI = 0 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
Here a lot of the different features indicate a non-neutral and probably damaging substitution, which begins with the complete different properties of the two amino acids. Furthermore the substitution matrices show low values and arginine is high conserved on position 123. Beside of SNAP all prediction tools forecast a bad substitution, however PolyPhen2 only indicates a possibly damaging mutation. Nevertheless SNAP itself has the lowest reliability index of 0. This substitution lies in a LOOP and a similar conformation of the two amino acids in the molecule can be seen in <xr id="R123I"/>. Altogether we predict that the substitution of arginine to isoleucine at position 123 of the PAH protein is non-neutral and might affect function.
Gln-20-Leu
| Properties | glutamine is neutral, polar and hydrophilic |
| leucine is aliphatic, neutral, non-polar and hydrophilic | |
| Substitution Matrices | low value → bad substitution |
| Structure | LOOP |
| PSSM | seems to be not conserved as the value for aspartic acid (17) is higher than for glutamine (14).
leucine has a value of 6. |
| SIFT | TOLERATED with a score of 0.90 |
| PolyPhen2 | benign with a score of 0.000 |
| SNAP | neutral, RI = 0 |
| MutationTaster | disease causing |
| ⇒ prediction: neutral | |
The exchange of glutamine to leucine at position 20 is predicted to be neutral, as there is much indication for this by most features. Only the substitution matrices indicate a bad substitution. Glutamine seems to be not conserved at this position, but flexible as there is a higher value for aspartic acid in PSSM. SIFT and PolyPhen2 both get high scores for being a neutral substitution, where SNAP has a very low reliability index. Only MutationTaster predicts it to be disease causing. Again the substitution can be found in a LOOP region.
Gln-172-His
<figure id="Q172H">
</figure>
| Properties | glutamine is neutral, polar and hydrophilic |
| histidine is positively charged, polar and hydrophobic | |
| Substitution Matrices | middle ranged → neutral substitution |
| Structure | LOOP |
| PSSM | ... |
| ... | |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.03. |
| PolyPhen2 | possibly damaging with a score of 0.705 / benign with a score of 0.170 |
| SNAP | neutral, RI = 4 |
| MutationTaster | disease causing |
| ⇒ prediction: neutral | |
Gly-103-Ser
| Properties | glycine is small, neutral, non-polar and hydrophilic |
| serine is small, neutral, polar and hydrophilic | |
| Substitution Matrices | middle ranged value → neutral substitution |
| Structure | HELIX |
| PSSM | glycine seems to be completely unconserved as every amino acid has the same value 5. |
| SIFT | TOLERATED with a score of 0.05 |
| PolyPhen2 | benign with a score of 0.003 / 0.006 |
| SNAP | neutral, RI = 6 |
| MutationTaster | disease causing |
| ⇒ prediction: neutral | |
Ile-421-Thr
<figure id="I421T">
</figure>
| Properties | isoleucine is aliphatic, neutral, non-polar and hydrophobic |
| threonine is small, neutral, polar and hydrophobic | |
| Substitution Matrices | low value for Blosum62, middle ranged value for PAM1/250 → bad or neutral substitution |
| Structure | STRAND |
| PSSM | isoleucine with a value of 32 seems to be not conserved as valine has a value of 41.
threonine has a value of 0. |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.00. |
| PolyPhen2 | possibly damaging with a score of 0.667 / probably damaging with a score of 0.913 |
| SNAP | non-neutral, RI = 2 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
Lys-341-Thr
<figure id="K341T">
</figure>
| Properties | lysine is positively charged, polar and hydrophobic |
| threonine is small, neutral, polar and hydrophobic | |
| Substitution Matrices | low value for Blosum62, middle ranged value for PAM1/250 → bad or neutral substitution |
| Structure | STRAND |
| PSSM | lysine seems to be conserved with a value of 39.
threonine has a value of 1. |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.00. |
| PolyPhen2 | probably damaging with a score of 1.000 / 0.996 |
| SNAP | non-neutral, RI = 3 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
Phe-392-Ser
<figure id="F392S">
</figure>
| Properties | phenylalanine is aromatic, neutral, non-polar and hydrophobic |
| serine is small, neutral, polar and hydrophilic | |
| Substitution Matrices | low value → bad substitution |
| Structure | HELIX |
| PSSM | phenylalanine seems to be highly conserved with a value of 49. Serine has a value of 0. |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.00. |
| PolyPhen2 | probably damaging with a score of 1.000 |
| SNAP | non-neutral, RI = 1 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
Pro-416-Gln
<figure id="P416Q">
</figure>
| Properties | proline is small, neutral, non-polar and hydrophilic |
| glutamine is neutral, polar and hydrophilic | |
| Substitution Matrices | low value → bad substitution |
| Structure | LOOP |
| PSSM | proline seems to be highly conserved with a value of 66. Glutamine has a value of 0. |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.00. |
| PolyPhen2 | probably damaging with a score of 0.996 / 0.985 |
| SNAP | non-neutral, RI = 1 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
Thr-266-Ala
<figure id="T266A">
</figure>
| Properties | threonine is small, neutral, polar and hydrophobic |
| alanine is small, non-polar, neutral and hydrophobic | |
| Substitution Matrices | middle ranged value &rarr neutral substitution |
| Structure | HELIX |
| PSSM | threonine seems to be conserved with a value of 43. Alanine has a value of 29. |
| SIFT | AFFECT PROTEIN FUNCTION with a score of 0.00. |
| PolyPhen2 | probably damaging with a score of 1.000 |
| SNAP | neutral, RI = 2 |
| MutationTaster | disease causing |
| ⇒ prediction: non-neutral | |
Mammalian Homologous Sequences
We used a normal BLAST search on the mammalian database of Uniprot and filter the results per hand for double entries. Altogether we found 22 homologues sequences (<xr id="IDs"/>). <figtable id="IDs">
| UniProt-IDs homologue to P00439 | |||||||
|---|---|---|---|---|---|---|---|
| H2Q6R0 | G3S964 | G1R3M2 | G7PJC2 | F7HMW9 | F7I717 | F7BKF9 | |
| F6XY00 | E2R366 | G1T8B6 | H2NIF5 | M3YKN3 | M9P0Q7 | H0WTI6 | |
| M3W9R1 | G3TIW0 | G1LIM6 | Q2KIH7 | G1P4I7 | P16331 | M9P0Y5 | |
</figtable>
For the creation of the multiple alignment we used clustalw. Nevertheless, only F6XY00 has a slightly different sequence. Like E2R366 F6XY00 is a homologue sequence of the human PAH in canis familiaris. On the positions of our substitutions the sequences are conserved.
Prediction
| SNP-Prediction | |||
|---|---|---|---|
| SNP | Prediction | Validation | |
| Ala259Val | neutral/non-neutral??? | HGMD | |
| Arg123Ile | non-neutral | ? | dbSNP |
| Gln20Leu | neutral | wrong | HGMD |
| Gln172His | neutral | ? | dbSNP |
| Gly103Ser | neutral | wrong | HGMD |
| Ile421Thr | non-neutral | ? | dbSNP |
| Lys341Thr | non-neutral | correct | HGMD |
| Phe392Ser | non-neutral | ? | dnSNP |
| Pro416Gln | non-neutral | correct | HGMD |
| Thr266Ala | non-neutral | correct | dbSNP |
References
<references/>