Difference between revisions of "Normal mode analysis HEXA"
From Bioinformatikpedia
(→Results) |
(→Results) |
||
| Line 186: | Line 186: | ||
=== Results === |
=== Results === |
||
| + | |||
| + | ==== 600K ==== |
||
=== Comparison to NOMAD-Ref === |
=== Comparison to NOMAD-Ref === |
||
Revision as of 12:30, 14 July 2011
Contents
Webnma
General information
Results
We analysed the five models with the lowest energy values. Webnma calculates fourteen different models with following energy values:
| Mode Index | Deformation Energy |
| 7 | 1041.88 |
| 8 | 1318.21 |
| 9 | 1738.30 |
| 10 | 2841.31 |
| 11 | 3135.09 |
| 12 | 4100.18 |
| 13 | 3911.06 |
| 14 | 5337.50 |
| 15 | 5741.69 |
| 16 | 6513.85 |
| 17 | 6081.05 |
| 18 | 6882.96 |
| 19 | 7514.14 |
| 20 | 7943.67 |
We took the first five models (7, 8, 9, 10, 11).
Here you can see the normalized squared atomic displacments of our models:
| model 7 | model 8 | model 9 | model 10 | model 11 |
Here you can see the motion of our analyses.
| model 7 | model 8 | model 9 | model 10 | model 11 |
Discussion
El Nemo
General information
Results
Discussion
Anisotropic Network Model web server
General information
Results
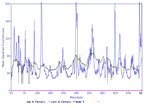
Here you can see the B-factor distribution of the real occuring B-factors (black) and the calulated B-factors (blue).
| model 1 | model 2 | model 3 | model 4 | model 5 |
In the next table, the motion of the protein of the different models is shown:
| model 1 | model 2 | model 3 | model 4 | model 5 |
Discussion
NOMAD-Ref
General information
Results
| model 1 | model 2 | model 3 | model 4 | model 5 |
| model 1 | model 2 | model 3 | model 4 | model 5 |