Difference between revisions of "Normal mode analysis (Phenylketonuria)"
(→WEBnm@) |
(→Mode 7) |
||
| Line 12: | Line 12: | ||
</figure><br clear=all> |
</figure><br clear=all> |
||
==== Mode 7 ==== |
==== Mode 7 ==== |
||
| − | <figure id="web_mode7"><small>[[File:PAH_1j8u_mode7.png|thumb|left|300px|'''<caption>'''...</caption>]]</small></figure> |
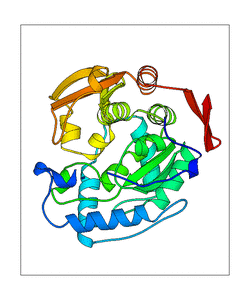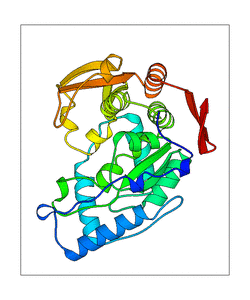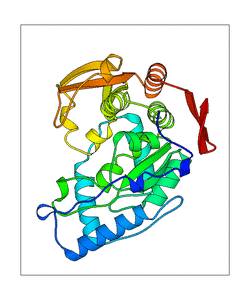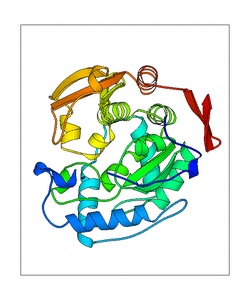
+ | <figure id="web_mode7"><small>[[File:PAH_1j8u_mode7.png|thumb|left|300px|'''<caption>'''The mode captured of WEBnm@. PDB structure is shown in transperent green, the ligands in red (H4B) and orange (Fe2). The blue doted lines represent the movement of the structure. </caption>]]</small></figure> |
| + | This mode seems to have a high movement |
||
| − | ... |
||
<br clear=all> |
<br clear=all> |
||
Revision as of 07:24, 30 July 2013
Page still under construction!!!
Contents
Summary
...
Normal Mode Analysis
Lab journal
For the analysis, we used the pdb structure 1J8U from the task before (Task 9 - Structure-based mutation analysis).
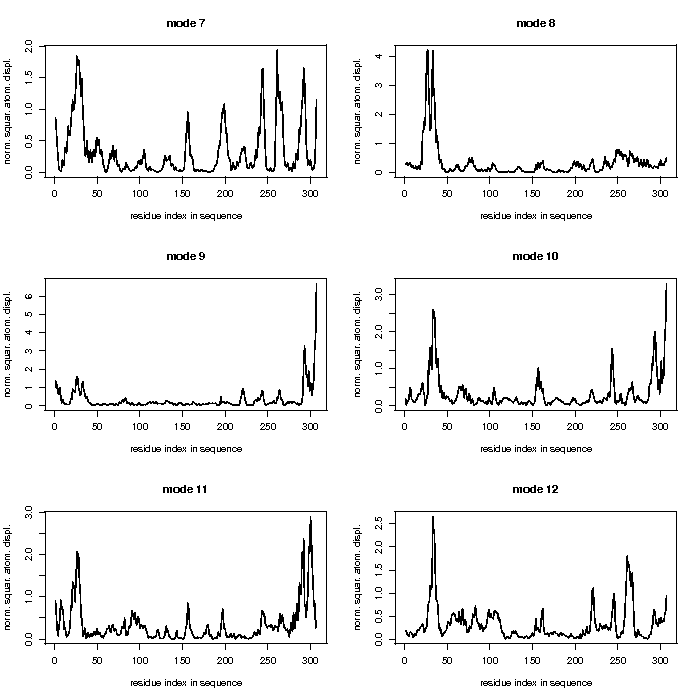
WEBnm@
<figure id="webMod">
</figure>
Mode 7
<figure id="web_mode7">
</figure>
This mode seems to have a high movement
Mode 8
<figure id="web_mode8">
</figure>
...
Mode 9
<figure id="web_mode9">
</figure>
...
Mode 10
<figure id="web_mode10">
</figure>
...
Mode 11
<figure id="web_mode11">
</figure>
...
Mode 12
<figure id="web_mode12">
</figure>
...
<figure id="correlation">
</figure>
elNémo
ElNémo computes models for the given protein 1J8U. Thereby it calculates 100 lowest-frequency modes and provides a lot of information about the modes with different parameters and visualizations:
- PDB-files with the conformations of the modes
- animations of the modes of three different sites
- CA-vari (calculates distance fluctuations between all C-alpha atoms)
- R2 residue mean square displacement of all C-alpha atoms
- Frequency and collectivity of the modes
<figtable id="frecol">
| First five modes | |||
|---|---|---|---|
| Mode | Frequency | Collectivity | |
| mode 7 | 1.00 | 0.5475 | |
| mode 8 | 1.02 | 0.4062 | |
| mode 9 | 1.36 | 0.2941 | |
| mode 10 | 1.43 | 0.0948 | |
| mode 11 | 1.60 | 0.3466 | |
</figtable>
- B-factor analysis for the correlation between observed and normal-mode-derived atomi displacement parameters:
- Correlation = 0.534 for 307 C-alpha atoms
- RMSD: only if two structures are used
Theoretical every number of normal mode perturbed models can be generated. However, a high number would take a lot of time. On default five normal modes will be computed by elNémo and can be increased up to 25. Additionally more normal modes can be calculated after the run. The analyses of the five modes with lowest frequency normal modes are shown below. For the calculation of the models all atoms are used.
Mode 7
<figure id="mode7_gif">
</figure>
<figure id="mode7_matrix">
</figure>
Mode 8
<figure id="mode8_gif">
</figure> <figure id="mode8_matrix">
</figure>
Mode 9
<figure id="mode9_gif">
</figure> <figure id="mode9_matrix">
</figure>
Mode 10
<figure id="mode10_gif">
</figure> <figure id="mode10_matrix">
</figure>
Mode 11
<figure id="mode11_gif">
</figure> <figure id="mode11_matrix">
</figure>
VMD
...
References
<references/>