Difference between revisions of "Structure-based mutation analysis TSD"
(→SCWRL and FoldX) |
(→Molecular Mechanics) |
||
| Line 192: | Line 192: | ||
|} |
|} |
||
</figtable> |
</figtable> |
||
| + | Since these are only the manually chosen side chain conformation, selected from the ones given by PyMOL, most of the analysis can be found in the sections below based on the predictions by SCWRL and FoldX. Here a few general things: |
||
| + | |||
| + | -- |
||
| + | |||
In the following, compare wild type (WT) and mutant structures. |
In the following, compare wild type (WT) and mutant structures. |
||
Revision as of 17:04, 25 June 2012
No citations, I don't read THAT many books :P - Have some music instead ;)
The journal of this task can be found here here.
Contents
Structure preparation
<figtable id="tbl:struct_comp">
| PDB-ID:Chain | Coverage | Resolution (Å) | R-value | R-free | pH |
|---|---|---|---|---|---|
| 2gjx:A | 23-529 | 2.8 | 0.270 | 0.288 | 5.5 |
| 2gk1:A | 23-529 | 3.25 | 0.277 | 0.322 | 5.5 |
</figtable>
There are two structures resolved for the HEXA_HUMAN reference sequence used for the course of this practical. The struture of this Uniprot entry is found in the alpha-chains of the PDB-IDs and 2gjx and 2gk1 both form the same publication <ref name="hexa_pdb_ref">Lemieux, M., Mark, B., & Cherney, M. (2006). Crystallographic Structure of Human beta-Hexosaminidase A: Interpretation of Tay-Sachs Mutations and Loss of GM2 Ganglioside Hydrolysis. Journal of molecular biology, 359(4), 913-29. doi:10.1016/j.jmb.2006.04.004</ref>. Unfortunately a 14 residue stretch towards the N-terminus of the protein (residues 75 to 88) is unresolved in both structures. However as previously shown the alpha subunit of Hex A consists of two domains. The N-terminal domain, Glyco_hydro_20b, is not involved in catalysis. Therefore, to evade problems with the missing backbone in the course of this task, the structure was truncated to contain only the C-terminal, catalytic domain, Glyco_hydro_20. In concordance with previous tasks and based on the experimental data in for both structures, shown in <xr id="tbl:struct_comp"/>, 2gjx was chosen as the reference structure. Details on the alteration of the structure according to the measures described above can be found in the journal.
Mutations
As only disease causing SNPs were assigned, the week before, some mutations had to be replaced. Additionally the reference PDB structure limited the possibilities, as only the second domain was retained for the following analysis. Thus an almost new set of SNPs was chosen: R178H, R178C, P182L, D207E, S293I, F434L, L451V, E482K, L484Q, E506D. Their position in the 3D-structure of HEXA is shown in <xr id="fig:snpsOnstr"/>.
<figure id="fig:snpsOnstr">
</figure>
The chosen mutations are displayed in the context of the remaining domain, the active site and important residues. For a detailed description of the important residues, see introduction . There are 5 mutations occurring in loop regions and 5 in helix elements. One intersection between mutations and important residues is at position 207 where there is a mutations from D to E. At the other important residue within the mutations, namely position 178 two mutants were chosen for the SNP set. Mutations of the important residues are expected to have a severe effect on the protein function.
For the new mutation set the biochemical properties are listed in <xr id="tab:biochem"/>. They have been assembled according to the sequence based mutation analysis.
<figtable id="tab:biochem">
| Mutation | Wildtype | Mutant | Grantham score | ||||
|---|---|---|---|---|---|---|---|
| Hydrophobicity | Volume | Charge | Hydrophobicity | Volume | Charge | ||
| R178H | -4.5 (polar) | 173.4 (bulky) | positive | -3.2 (polar) | 153.2 (bulky) | neutral | 29 |
| R178C | -4.5 (polar) | 173.4 (bulky) | positive | 2.5 (polar) | 108.5 (small) | neutral | 180 |
| P182L | -1.6 (nonpolar) | 112.7 (small) | neutral | 3.8 (nonpolar) | 166.7 (bulky) | neutral | 98 |
| D207E | -3.5 (polar) | 111.1 (small) | negative | -3.5 (polar) | 138.4 (bulky) | negative | 45 |
| S2931 | -0.8 (polar) | 89.0 (tiny) | neutral | 4.5 (nonpolar) | 166.7 (bulky) | neutral | 142 |
| S434L | 2.8 (nonpolar) | 189.9 (bulky) | neutral | 3.8 (nonpolar) | 166.7 (bulky) | neutral | 22 |
| L451V | 3.8 (nonpolar) | 166.7 (bulky) | neutral | 4.2 (nonpolar) | 140.0 (small) | neutral | 32 |
| E482K | -3.5 (polar) | 138.4 (bulky) | negative | -3.9 (polar) | 168.6 (bulky) | positive | 56 |
| 484 | 3.8 (nonpolar) | 166.7 (bulky) | neutral | -3.5 (polar) | 143.8 (bulky) | neutral | 113 |
| E506D | -3.5 (polar) | 138.4 (bulky) | negative | -3.5 (polar) | 111.1 (small) | negative | 45 |
</figtable>
Map all the 10 mutations onto the crystal structure. Color the mutants differently than the rest of the protein and create a snapshot for the wiki. If applicable find out whether the mutations are close to the active site, a binding interface or other important functional sites. Visualize this and describe it properly.
Molecular Mechanics
<figtable id="tbl:pymolstruc">
</figtable> Since these are only the manually chosen side chain conformation, selected from the ones given by PyMOL, most of the analysis can be found in the sections below based on the predictions by SCWRL and FoldX. Here a few general things:
--
In the following, compare wild type (WT) and mutant structures.
Now that you should have a clear idea of the WT and mutant proteins we will try to calculate some energies. Always calculate the energy for the wild type and mutants – then substract/compare.
SCWRL and FoldX
<figtable id="tbl:pymolstruc">
</figtable>
R178H/C
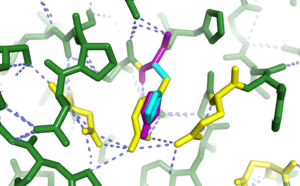
R178 is one of the important residues. As can be seen in the according figure in <xr id="tbl:pymolstruc"/> it could form hydrogen bonds with the neighbouring residues D207 and E462. In addition it seems reasonable that these two neighbouring residues with the negative charges somehow coordinate the positive charges placed between them. More importantly R178 has already been shown to form a hydrogen bond with the ligand.
The mutant amino acid histidine, depending on the pH value, can also carry a positive charge. The reported optimum pH value for Hex A is around 4.4, in these conditions, histidine is sure to contain a positive charge <ref name="phforhexa">Grebner,E.E. et al. (1986) Two abnormalities of hexosaminidase A in clinically normal individuals. American journal of human genetics, 38, 505-14.</ref><ref name="histcharge">when is his pos charged, alberts TODO</ref>. However the larger problem under is that under these conditions the formation of a hydrogen bond with the ligand is very unlikely. This is reinforced by the recessed positioning of the residue relative to the active site. Even for the case of SCWRL (purple), where the hydrogen is included (and the ring flipped compared to FoldX), the distance could already be too long for coordination of the ligand. There are slight clashes of both the SCWRL as well as the FoldX structures with D175 however these are very minor, fixable and clearly not the main problem here.
For the mutation to cysteine, the problem is much more pronounced. There is no doubt that formation of the hydrogen bond to the ligand is not possible anymore due to the increase in distance. Clashes do not exist, but would actually be hard to create in any case.
In conclusion the modelling of both mutation side chains is mostly as good as possible and shows only minor differences between the two methods. The orientation of the histidine ring is however more convincing in the SCWRL result.
P182L
D207E
S293I
F434L
L451V
E482K
L484Q
E506D
Investigate the local hydrogen-bonding network using pymol[1] – also check for potential clashes (when sidechains are too close to each other). Are you introducing hydrophilics to the core or hydrophobics to the protein surface? Are there any holes introduced to the protein due to the mutations? Foreach of the mutations also a new structure will be created. Note down all of the energies, but also use these structures in the next steps.
Compare the scwrl and foldx structures in Pymol and superimpose them. What are the differences?
Minimise
What happens regarding the energy?
Gromacs
Analyze the minimization of the system with the following command: g_energy -f FILE.edr -o energy_1.xvg. Do the analysis for Bond, Angle and Potential. The xvg graphs can be viewed with xmgrace and in the print settings you can choose eps output, the print and convert to pdf.