Difference between revisions of "Task 10: Molecular Dynamics Analysis"
(→HYDROGEN BONDS) |
(→Cluster Analysis) |
||
| Line 616: | Line 616: | ||
==== Cluster Analysis ==== |
==== Cluster Analysis ==== |
||
| + | |||
| + | |||
| + | <code> |
||
| + | //when asked for input select 6 and 6 again |
||
| + | g_cluster -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -dm /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-matrix.xpm -dist /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-distribution.xvg -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/clusters.xpm -sz /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-sizes.xvg -tr /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-transitions.xpm -ntr /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-transitions.xvg -clid /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-id-over-time.xvg -cl /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/clusters.pdb -cutoff 0.1 -method gromos -dt 10 |
||
| + | |||
| + | </code> |
||
==P281L== |
==P281L== |
||
Revision as of 08:23, 16 September 2011
In this task we are going to analyze the results of the molecular dynamics simulations of task 8. A detailed task description can be found here. The analysis focuses on this tutorial.
Contents
- 1 Native
- 2 P281L
- 3 R408W
Native
A BRIEF CHECK OF RESULTS
In order to verify that our simulation ran successfully we used the command line tool gmxcheck. we executed it as follows for our .xtc file:
gmxcheck -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.xtc
How many frames are in the trajectory file and what is the time resolution?
We observed 2001 frames with a time resolution of 5ps.
How long did the simulation run in real time (hours), what was the simulation speed (ns/day) and how many years would the simulation take to reach a second?
The simulation ran 4h00:39 and had a simulation speed of 59.835 ns/day. To calculate 1 second we would need (1 / (59 * (10^(-9)))) / 365 = 46 436.0344 years
Which contribution to the potential energy accounts for most of the calculations?
- potential energy: -4.57312e+05 kJ/mol
VISUALIZATION OF RESULTS
We extracted 1000 frames from the trajectory (-dt 10), leaving out the water (selected Protein when asked for a selection). Moreover, we will remove the jumps over the boundaries and make a continuous trajectory (-pbc nojump):
trjconv -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.xtc -o protein.pdb -pbc nojump -dt 10
After that we opened the generated protein.pdb file with pymol. Here we changed the coloring to spectrum by typing the following to the pymol command line:
spectrum
In a next step we enabled the the visualization of the cell with this command:
show cell
In order to remove the tumbeling and wiggeling motion of our protein we used the command intra_fit since we are only interested in the internal motions of the protein:
intra_fit protein
The results of these actions can be seen in figure 1 and 2. Figure one shows our WT protein in line view which makes it able to see the motion of the side chains. Figure to shows the protein in cartoon view to see the overall movement of the secondary structure elements.
QUALITY ASSURANCE
CONVERGENCE OF ENERGY TERMS
In this part of quality assurance we analyzed different metrics of our MD simulation by creating plots from our *.edr file. We created plots for the temperature, pressure, energy, volume, density, box, coulomb and van der waals values of our MD simulation by using the tool g_energy as follows:
//calculating temperature enter then "12 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/temperature.xvg
//calculating pressure enter then "13 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/pressure.xvg
//calculating energy (potential, kinetic and total) enter then "9\n10\n11 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/energy.xvg
//calculating volume enter then "18 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/volume.xvg
//calculating density enter then "19 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/density.xvg
//calculating box enter then "15\n16\n17 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/box.xvg
//calculating coulomb enter then "48\n50 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/coulomb-inter.xvg
//calculating van der waals enter then "49\n51 0"
g_energy -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.edr -o ../../../Dropbox/Studium/3_semester/master_praktikum/task10/vanderwaals-inter.xvg
Temperature Over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Temperature" | 297.886 | 0.007 | 1.57824 | "-0.0105059 (K)" |
In figure 3 we can see that the temperature stays quite the same over the whole simulation which might be interpreted as that our system has reached its stable temperature for simulation.
Pressure over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Pressure" | 1.02313 | 0.023 | 133.659 | "-0.0928554 (bar)" |
Energy over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Potential" | -457312 | 61 | 640.502 | " -423.792 (kJ/mol)" |
| "Kinetic En." | 81855.1 | 1.9 | 433.677 | " -2.88684 (kJ/mol)" |
| "Total Energy" | -375457 | 61 | 784.246 | " -426.678 (kJ/mol)" |
Volume over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Volume" | 356.629 | 0.041 | 0.397685 | " -0.151337 (nm^3)" |
Density over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Density" | 1021.28 | 0.12 | 1.13884 | "0.433467 (kg/m^3)" |
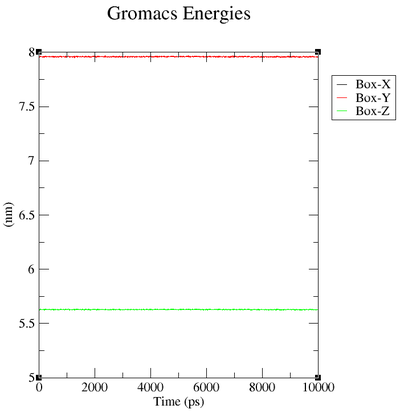
Box over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Box-X" | 7.95996 | " 0.0003" | 0.00295875 | "-0.00112603 (nm)" |
| "Box-Y" | 7.95996 | " 0.0003" | 0.00295875 | "-0.00112603 (nm)" |
| "Box-Z" | 5.62854 | 0.00021 | 0.00209216 | "-0.000796226 (nm)" |
Coulomb over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Coul-SR:Protein-non-Protein" | " -20429.9" | "110" | "425.888" | "-561.713 (kJ/mol)" |
| "Coul-14:Protein-non-Protein" | " 0" | " 0" | " 0" | " 0 (kJ/mol)" |
Van der Waals over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "LJ-SR:Protein-non-Protein" | "-2172.28" | "15" | "136.956" | "-103.12 (kJ/mol)" |
| "LJ-14:Protein-non-Protein" | " 0" | " 0" | " 0" | " 0 (kJ/mol)" |
Questions
1. What is the average temperature and what is the heat capacity of the system?
The average temperature of our system is 297.886 K and the heat capacity ranges from 303 to 292. (values taken from figure 3).
2. What are the terms plotted in the files energy.xvg and box.xvg
In energy.xvg we plot the energy values of kinetic energy, potential energy and total energy over the time of our simulation.
In box.xvg we plot the size of the box around our protein which is given in nm.
3. Estimate the plateau values for the pressure, the volume and the density.
We consider the plateau value as two values, the upper and lower plateau. These plateaus represent the maximum and minimum values of our given plots.
Pressure:
- upper plateau:450 bar
- lower plateau: -450 bar
Volume:
- upper plateau: 357.9 nm^3
- lower plateau: 355.8 nm^3
Density:
- upper plateau: 1026 kg/m^3
- lower plateau: 1017 kg/m^3
4. What are the terms plotted in the files coulomb-inter.xvg and vanderwaals-inter.xvg
The coloumb and van der waals energys of our WT over time.
MINIMUM DISTANCES BETWEEN PERIODIC IMAGES
We ran gromacs with the following command:
//when asked for group selection we selected group 1
g_mindist -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -od /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/minimal-periodic-distance.xvg -pi
And produced the plot seen in figure 11.
In a next step we produced a plot which only uses the C-alpha atoms to calculate the minimum distance between periodic images. For this we used the same command as before. Although, we selected group 3 this time. The result can be seen in figure 12.
Questions
What was the minimal distance between periodic images and at what time did that occur?
The shortest periodic distance is 1.73442 (nm) at time 2720 (ps), between atoms 563 and 5314.
What happens if the minimal distance becomes shorter than the cut-off distance used for electrostatic interactions? Is it the case in your simulations?
If the distance would become shorter than the cut-off distance used for electrostatic interactions our energy would dramatically increase. This did not happen for our simulation so we can assume that our measured value of 1.73 nm is still higher than the cutoff.
Run now g_mindist on the C-alpha group, does it change the results? What does is mean for your system?
With C-alpha group we got a shortest periodic distance of 2.4249 (nm) at time 6805 (ps), between atoms 589 and 5297. This is an increase of the distance since we only consider C-alpha atoms and no side chains as in our previous calculation.
ROOT MEAN SQUARE FLUCTUATIONS
In this part of the analysis we are going to have a look at the root mean square fluctuations. By analyzing this value for our structure we might be able to figure out which parts of our protein are more flexible than others.
To do so we executed the following command:
g_rmsf -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsf-per-residue.xvg -ox /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/average.pdb -oq /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/bfactors.pdb -res
Which gives us three result files:
- rmsf-per-residue.xvg : this file contains a plot over the whole structure length of the RMSF values
- average.pdb : is the average conformation of the structure
- bfactors.pdb: assigned b factor values to the residues which gives us information about the flexibility
Questions
Indicate the start and end residue for the most flexible regions and the maximum amplitudes.
The most flexible region in our structure is from position 131 to 152 which is a loop connecting two alpha-helices. This can be observed in the plot in figure 13 as well as in the 3D view of our protein in figure 14. In general we can say that most of the parts are rather rigid, especially the residues which are located within the core. We observe slightly more flexibility in some alpha-helices which are located at the surface of the protein. Such as the residues located at position 197, 208, 215 and 216. There are also some beta-sheets which contain slightly flexible residues such as residues at position 413 and 420.
CONVERGENCE OF RMSD
It is adviced in the tutorial to remove the jumps of our protein and bring it back to the middle. This is done with the following command:
trjconv -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.xtc -o 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -pbc nojump
Then we calculated the RMSD for the whole protein and a second time only for the backbone with the following command:
//whole protein: select 1 and 1 again
g_rms -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-all-atom-vs-start.xvg
//backbone only: select 4 and 4 again
g_rms -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-backbone-vs-start.xvg
After that we calculated the RMSD for the average protein structure with the following commands, once for the whole protein and once for the backbone only:
trjconv -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/protein.xtc
g_rms -f /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/protein.xtc -s /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/average.pdb -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-all-atom-vs-average.xvg
g_rms -f /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/protein.xtc -s /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/average.pdb -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-backbone-vs-average.xvg
We received then the following results:
Questions
If observed, at what time and value does the RMSD reach a plateau?
We observed in the figures (16-17, against start structure) that the plateau is reached after 5000ps with the following approximated values:
- All atom vs start: 0.225
- Backbone vs start: 0.175
For the average structures we observed that the plateau is reached earlier after 300ps with the following approximated values:
- All atom vs average: 0.16
- Backbone vs average: 0.125
Briefly discuss two differences between the graphs against the starting structure and against the average structure. Which is a better measure for convergence?
As we can see the plateau is reached earlier in the average structure than in the start structure. The reason for this might be that the average structure is closer to the equilibrium than the start structure. Hence, also the RMSD values are lower here.
Since the average structures converges earlier than the starting structure we think that this plots are the better ones to judge whether convergence has been reached or not.
CONVERGENCE OF RADIUS OF GYRATION
To calculate the radius of gyration we executed the following command:
g_gyrate -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/radius-of-gyration.xvg
And received the following plot :
Questions
At what time and value does the radius of gyration converge?
The radius of gyration fluctuates between 1.90 to 1.95 nm. The value for RGz is fluctuating between 1.55 to 1.70, RGx is fluctuating between 1.45 to 1.70 and RGy is fluctuating between 1.45 to 1.55.
STRUCTURAL ANALYSIS: PROPERTIES DERIVED FROM CONFIGURATIONS
SOLVENT ACCESSIBLE SURFACE AREA
In this part of our analysis we are checking the solvent accessible area of our protein over time. To calculate these values we executed the following command:
// select 1 and 1 again when asked for group
g_sas -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/solvent-accessible-surface.xvg -oa /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/atomic-sas.xvg -or /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/residue-sas.xvg
We received the following 3 output plots:
HYDROGEN BONDS
In this part of our analysis we are looking at inter H-bonds within our protein and at H-bonds from our protein to the surrounding solvent. To do so we executed the following command twice:
// first time with 1 and 1 again to calculate inter H-bonds
// second time with 1 and 12 to calculate H-bonds from the protein to the surrounding solvent
g_hbond -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -num /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/hydrogen-bonds-protein-water.xvg
We received the following results seen in the figures below:
Questions
Discuss the relation between the number of hydrogen bonds for both cases and the fluctuations in each.
SALT BRIDGES
This part of the analysis could not be finished yet because of a lack of computational power.
RAMACHANDRAN (PHI/PSI) PLOTS
In this part of the analysis we have a look at the phi/psi angles in our protein. To do the calculations we executed the following command:
g_rama -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/ramachandran.xvg
We received the following plot:
ANALYSIS OF DYNAMICS AND TIME-AVERAGED PROPERTIES
ROOT MEAN SQUARE DEVIATIONS AGAIN
In this part of the analysis we calculate the RMSD of our structure versus our structure over time. To do so we executed the following two commands:
//RMSD calculation
g_rms -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -f2 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -m rmsd-matrix.xpm -dt 10
//coloring of plot
xpm2ps -f rmsd-matrix.xpm -o rmsd-matrix.eps -rainbow blue
Finally we received the following plot:
Questions
What is interesting by choosing the group "Mainchain+Cb" for this analysis? Think about the different proteins used for this practical.
How many transitions do you see?
What can you conclude from this analysis? Could you expect such a result, justify?
Cluster Analysis
//when asked for input select 6 and 6 again
g_cluster -s 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md.tpr -f 1J8U_nosol_after_SCWRL_no_h_merged_crystal_water_md_nojump.xtc -dm /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-matrix.xpm -dist /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/rmsd-distribution.xvg -o /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/clusters.xpm -sz /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-sizes.xvg -tr /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-transitions.xpm -ntr /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-transitions.xvg -clid /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/cluster-id-over-time.xvg -cl /home/student/Dropbox/Studium/3_semester/master_praktikum/task10/clusters.pdb -cutoff 0.1 -method gromos -dt 10
P281L
A BRIEF CHECK OF RESULTS
In order to verify that our simulation ran successfully we used the command line tool gmxcheck. we executed it as follows for our .xtc file:
gmxcheck -f P281L_md.xtc
How many frames are in the trajectory file and what is the time resolution?
We observed 2001 frames with a time resolution of 5ps.
How long did the simulation run in real time (hours), what was the simulation speed (ns/day) and how many years would the simulation take to reach a second?
The simulation ran 3h14 and had a simulation speed of 73.133 ns/day. To calculate 1 second we would need 0.328*10^9/(365*24) = 37442.92 years
Which contribution to the potential energy accounts for most of the calculations?
- potential energy: -4.57346e+05 kJ/mol
- greatest contribution by Coulomb: -4.45954e+05 kJ/mol
VISUALIZATION OF RESULTS
We extracted 1000 frames from the trajectory (-dt 10), leaving out the water (selected Protein when asked for a selection). Moreover, we will remove the jumps over the boundaries and make a continuous trajectory (-pbc nojump):
trjconv -s P281L_md.tpr -f P281L_md.xtc -o protein.pdb -pbc nojump -dt 10
After that we opened the generated protein.pdb file with pymol.
QUALITY ASSURANCE
CONVERGENCE OF ENERGY TERMS
In this part of quality assurance we analyze different metrics of our MD simulation by creating plots from our *.edr file. We created plots for for pressure, temperature, potential and total energy of our MD simulation.
Temperature Over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Temperature" | 297.887 | 0.0054 | 1.58026 | 0.0204823 |
In figure 3 we can see that the temperature stays quite the same over the whole simulation which might be interpreted as that our system has reached its stable temperature for simulation.
Pressure over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Pressure" | 1.01427 | 0.011 | 133.436 | -0.0559964 |
Total Energy over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Total Energy" | -375514 | 53 | 783.088 | -307.507 |
Potential Energy over Time
| "Energy" | "Average" | "Err.Est." | "RMSD" | "Tot-Drift" |
| "Total Energy" | -457346 | 54 | 638.456 | -313.132 |
MINIMUM DISTANCES BETWEEN PERIODIC IMAGES
We ran gromacs with the following command:
//when asked for group selection we selected group 1
g_mindist -f P281L_md.xtc -s P281L_md.tpr -od minimal-periodic-distance.xvg -pi
And produced the plot seen in figure 11.
In a next step we produced a plot which only uses the C-alpha atoms to calculate the minimum distance between periodic images. For this we used the same command as before. Although, we selected group 3 this time. The result can be seen in figure 12.
Questions
What was the minimal distance between periodic images and at what time did that occur?
The shortest periodic distance is 1.9083 (nm) at time 55 (ps), between atoms 1198 and 5323.
What happens if the minimal distance becomes shorter than the cut-off distance used for electrostatic interactions? Is it the case in your simulations?
If the distance would become shorter than the cut-off distance used for electrostatic interactions our energy would dramatically increase. This did not happen for our simulation so we can assume that our measured value of 1.9083 nm is still higher than the cutoff.
Run now g_mindist on the C-alpha group, does it change the results? What does is mean for your system?
With C-alpha group we got a shortest periodic distance is 2.75268 (nm) at time 2320 (ps), between atoms 2715 and 5306. This is an increase of the distance since we only consider C-alpha atoms and no side chains as in our previous calculation.
ROOT MEAN SQUARE FLUCTUATIONS
In this part of the analysis we are going to have a look at the root mean square fluctuations. By analyzing this value for our structure we might be able to figure out which parts of our protein are more flexible than others.
Questions
Indicate the start and end residue for the most flexible regions and the maximum amplitudes.
CONVERGENCE OF RADIUS OF GYRATION
Questions
At what time and value does the radius of gyration converge?
STRUCTURAL ANALYSIS: PROPERTIES DERIVED FROM CONFIGURATIONS
SOLVENT ACCESSIBLE SURFACE AREA
HYDROGEN BONDS
In this part of our analysis we are looking at inter H-bonds within our protein and at H-bonds from our protein to the surrounding solvent. To do so we executed the following command twice: