Difference between revisions of "Structure-based mutation analysis ARSA"
(→FoldX) |
(→Visualization with Pymol) |
||
| Line 20: | Line 20: | ||
| Asp-Asn |
| Asp-Asn |
||
| 29 |
| 29 |
||
| − | | [[File:arsa29.png | 200px |
+ | | [[File:arsa29.png | 200px]] |
| The mutation is located at the position of a metal-binding site. |
| The mutation is located at the position of a metal-binding site. |
||
| harmful |
| harmful |
||
| Line 27: | Line 27: | ||
| Pro - Ala |
| Pro - Ala |
||
| 136 |
| 136 |
||
| − | | [[File:arsa136.png | 200px |
+ | | [[File:arsa136.png | 200px ]] |
| The mutation is located near the active site and a substrate binding site in sequence as well as in structure. |
| The mutation is located near the active site and a substrate binding site in sequence as well as in structure. |
||
| harmful |
| harmful |
||
| Line 34: | Line 34: | ||
| Gln-His |
| Gln-His |
||
| 153 |
| 153 |
||
| − | | [[File:arsa153.png | 200px |
+ | | [[File:arsa153.png | 200px ]] |
| The mutation is located near the active site and a substrate binding site in sequence as well as in structure. |
| The mutation is located near the active site and a substrate binding site in sequence as well as in structure. |
||
| harmful |
| harmful |
||
| Line 41: | Line 41: | ||
| Trp-Cys |
| Trp-Cys |
||
| 193 |
| 193 |
||
| − | | [[File:arsa193.png | 200px |
+ | | [[File:arsa193.png | 200px ]] |
| The mutation is at moderate distance to all important functional sites of the protein. |
| The mutation is at moderate distance to all important functional sites of the protein. |
||
| neutral |
| neutral |
||
| Line 48: | Line 48: | ||
| Thr-Met |
| Thr-Met |
||
| 274 |
| 274 |
||
| − | | [[File:arsa274.png | 200px |
+ | | [[File:arsa274.png | 200px ]] |
| The mutation is located very distant from the active site and a substrate binding site in sequence as well as in structure. <br> It is located within a beta sheet. |
| The mutation is located very distant from the active site and a substrate binding site in sequence as well as in structure. <br> It is located within a beta sheet. |
||
| harmful |
| harmful |
||
| Line 55: | Line 55: | ||
| Phe -Val |
| Phe -Val |
||
| 356 |
| 356 |
||
| − | | [[File:arsa356.png | 200px |
+ | | [[File:arsa356.png | 200px ]] |
| The mutation is at moderate distance to all important functional sites of the protein. |
| The mutation is at moderate distance to all important functional sites of the protein. |
||
| neutral |
| neutral |
||
| Line 62: | Line 62: | ||
| Thr-Ile |
| Thr-Ile |
||
| 409 |
| 409 |
||
| − | | [[File:arsa409.png | 200px |
+ | | [[File:arsa409.png | 200px ]] |
| The mutation is not close to important functional sites. |
| The mutation is not close to important functional sites. |
||
| harmful |
| harmful |
||
| Line 69: | Line 69: | ||
| Asn-Ser |
| Asn-Ser |
||
| 440 |
| 440 |
||
| − | | [[File:arsa440.png | 200px |
+ | | [[File:arsa440.png | 200px ]] |
| The mutation is very distant from all functional sites. |
| The mutation is very distant from all functional sites. |
||
| neutral |
| neutral |
||
| Line 76: | Line 76: | ||
| Cys-Gly |
| Cys-Gly |
||
| 489 |
| 489 |
||
| − | | [[File:arsa489.png | 200px |
+ | | [[File:arsa489.png | 200px ]] |
| The mutation is very distant from all functional sites. |
| The mutation is very distant from all functional sites. |
||
| harmful |
| harmful |
||
| Line 83: | Line 83: | ||
| Arg-His |
| Arg-His |
||
| 496 |
| 496 |
||
| − | | [[File:arsa496.png | 200px |
+ | | [[File:arsa496.png | 200px ]] |
| The mutation is very distant from all functional sites. |
| The mutation is very distant from all functional sites. |
||
| harmful |
| harmful |
||
Revision as of 16:41, 30 June 2011
Preparation
Visualization with Pymol
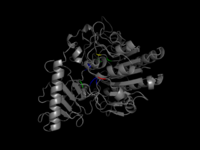
The following image shows a pymol visualization of Arylsulfatase A, together with all known active and binding sites.
One can see, that the mutations are spread out through the protein. Some lie near functional sites, other are very distant from them. The table below shows again Pymol visualizations of all mutations, but each seperately. With this, we want to try to derive investigate the correlation of location of the mutation with respect to functional sites and its effect on the protein function.
Above visualizations indicate, that mutations near functional important sites of the protein are likely to cause a harmful effect. However, for distant mutations no trend can be observed.
SCRWL
First, we extracted the amino acid sequence from our pdb file and converted it to lower case.
repairPDB arsa_model.pdb -seq > arsa.model.seq
tr '[:upper:]' '[:lower:]' < arsa.model.seq > arsa.model.lower.seq
Next, we included the individual mutations as capital letters in seperate files and executed scwrl with the following command:
scwrl cmd
We also ran SCWRL on the wild type (wt) structure in order to make it comparable to energy predictions by other programs. The minimal energy of the graph for the wt is 415.134.
FoldX
wt: 510.88
| Nr. | mutation | position | Minimal energy | Energy(mutant)/Energy(wt) | |
| 1 | Asp-Asn | 29 | 496.88 | 0.9725963 | |
| 2 | Pro - Ala | 136 | 493.81 | 0.966587 | |
| 3 | Gln-His | 153 | 493.90 | 0.9667632 | |
| 4 | Trp-Cys | 193 | 496.23 | 0.971324 | |
| 5 | Thr-Met | 274 | 503.34 | 0.9852412 | |
| 6 | Phe -Val | 356 | 495.39 | 0.9696798 | |
| 7 | Thr-Ile | 409 | 495.45 | 0.9697972 | |
| 8 | Asn-Ser | 440 | 496.79 | 0.9724201 | |
| 9 | Cys-Gly | 489 | 495.87 | 0.9706193 | |
| 10 | Arg-His | 496 | 498.75 | 0.9762567 |
FoldX
wt: -3839.492677
| Nr. | mutation | position | Free energy | Energy(mutant)/Energy(wt) |
| 1 | Asp-Asn | 29 | -4174.487222 | 1.087250 |
| 2 | Pro - Ala | 136 | -4164.523707 | 1.084655 |
| 3 | Gln-His | 153 | -4109.12924 | 1.070227 |
| 4 | Trp-Cys | 193 | -4169.617285 | 1.085981 |
| 5 | Thr-Met | 274 | -4065.730562 | 1.058924 |
| 6 | Phe -Val | 356 | -4109.021939 | 1.070199 |
| 7 | Thr-Ile | 409 | -4121.130656 | 1.073353 |
| 8 | Asn-Ser | 440 | -4120.275589 | 1.073130 |
| 9 | Cys-Gly | 489 | -4127.969116 | 1.075134 |
| 10 | Arg-His | 496 | -4120.151911 | 1.073098 |