Difference between revisions of "Hemochromatosis: Sequence based predictions"
| Line 1: | Line 1: | ||
| − | [[Lab |
+ | [[Lab Journall Task 3 Hemo||Lab Journa]] |
==Secondary Structure== |
==Secondary Structure== |
||
Revision as of 13:54, 26 August 2013
Contents
Secondary Structure
In this task, secondary structure of proteins is predicted using the programs reProf and PsiPred. The results are then compared to the DSSP[1] secondary structure assignments from corresponding crystal structures in the PDB. Thus, the following crystal structures were selected for comparison. The first priority for selection was to get the protein in its native state and not bound to another molecule. The other criteria were the quality of the structure and the alignment to the protein sequence(having one continous segment).
| Uniprot | PDB | |||||
|---|---|---|---|---|---|---|
| UID | Name | Length | ID | Resolution [A] | Chain | Length |
| Q30201 | Hereditary hemochromatosis protein | 348 | A16Z | 2.60 | A | 275 |
| P10775 | Ribonuclease inhibitor | 456 | 2BNH | 2.30 | A | 457 |
| Q9X0E6 | Divalent-cation tolerance protein CutA | 101 | 1VHF | 1.54 | A | 113 |
| Q08209 | CAM-PRP catalytic subunit | 521 | 1M63 | 2.80 | A | 372 |
In the next step, the output of the prediction programs and the DSSP assignments have to be made comparable. DSSP assigns 8 different classes of secondary structure, whereas reProf and PsiPred only predict helix(H), sheet(E) and loop(L or C). Therefore, H and G are mapped to H, E to E and all other DSSP classes to C.
To assess the quality of the prediction, the class specific accuracy, coverage and F1-measure were used along with the Q3 and SOV3<ref>Zemla. A, et al.;A Modified Definition of Sov, a Segment-Based Measure for Protein Secondary Structure Prediction Assessment; PROTEINS 34:220–223 (1999)</ref> measures. The Q3 is defined as follows:
<math> Q_3 = \frac{\text{correct predictions}}{ \text{all predictions}} = \frac{TP + TN}{TP + FP + TN + FN } } </math>
Reprof takes as input either sequences or PSSMs. Therefore PSSMs were generated by querying the HFE(Q30201) sequence against the big_80, SwissProt and PDB databases. The tool of choice for this was PsiBlast with standard parameters(2 iterations and e-value cutoff of 0.002).
| Prediction methods | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Reprof + Sequence | Reprof + Big80 | Reprof + SwissProt | Reprof + PDB | Psipred | |||||||||||
| Q3 | 0.76 | 0.81 | 0.86 | 0.84 | 0.84 | ||||||||||
| SOV3 | 0.66 | 0.75 | 0.84 | 0.84 | 0.73 | ||||||||||
| Acc | Cov | F1 | Acc | Cov | F1 | Acc | Cov | F1 | Acc | Cov | F1 | Acc | Cov | F1 | |
| H | 0.63 | 0.33 | 0.44 | 0.74 | 0.48 | 0.59 | 0.84 | 0.65 | 0.74 | 0.85 | 0.52 | 0.64 | 0.98 | 0.77 | 0.86 |
| C | 0.57 | 0.69 | 0.63 | 0.63 | 0.75 | 0.68 | 0.68 | 0.81 | 0.74 | 0.64 | 0.83 | 0.72 | 0.61 | 0.95 | 0.74 |
| E | 0.79 | 0.63 | 0.70 | 0.81 | 0.84 | 0.83 | 0.89 | 0.86 | 0.87 | 0.88 | 0.85 | 0.86 | 0.94 | 0.55 | 0.69 |
The deciding criteria for the reprof method choice were the Q3, SOV3 and F1-measures. For the Q3 and the SOV3, the SwissProt PSSM performed best and the F1-measures were also reasonable. Therefore, this method was chosen for all future predictions.
| Protein | Method | Q3 | SOV3 |
|---|---|---|---|
| Q08209 | ReProf | 0.83 | 0.78 |
| Psipred | 0.87 | 0.79 | |
| Q30201 | ReProf | 0.86 | 0.84 |
| Psipred | 0.84 | 0.73 | |
| P10775 | ReProf | 0.91 | 0.93 |
| Psipred | 0.93 | 0.94 | |
| Q9X0E6 | ReProf | 0.75 | 0.65 |
| Psipred | 0.89 | 0.86 |
While SOV3 and Q3 do not always agree which of the two methods performs better, they do so most of the time. Nevertheless, it was decided that SOV3 is to be trusted more than Q3, because it takes into account the per segment accuracy rather than just the per residue accuracy. Thus, PsiPred outperforms reprof in 3 out of 4 cases. It is also notable, that the SOV3 values range from 0.65 for Q9X0E6(101 residues) to 0.94 for P10775(456 residues). So the performance of the method depends on the length of the protein and the protein itself, but PsiPred performed best overall.
Disorder
| UID | DisProt ID | SeqID | E-value | Method |
|---|---|---|---|---|
| Q08209 | DP00092 | - | - | direct match |
| Q30201 | DP00670 | 0.29 | 2e-30 | psiblast |
| P10775 | DP00465 | 0.4 | 7e-30 | psiblast |
| Q9X0E6 | DP00564 | 0.25 | 0.36 | smith-waterman |
Transmembrane Helices
| Uniprot | PDB | |||||
|---|---|---|---|---|---|---|
| UID | Name | Length | ID | Resolution [A] | Chain | Length |
| Q30201 | Hereditary hemochromatosis protein | 348 | A16Z | 2.60 | A | 275 |
| Q9YDF8 | Voltage-gated potassium channel | 295 | 1ORQ | 3.20 | C | 223 |
| P35462 | D(3) dopamine receptor | 400 | 3PBL | 2.89 | A | 481 |
| P47863 | Aquaporin-4 | 323 | 2D57 | 3.20 | A | 301 |
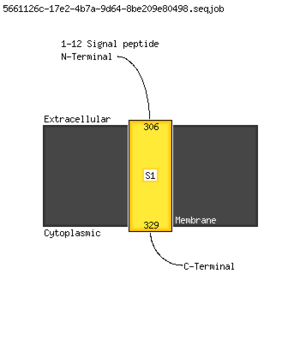
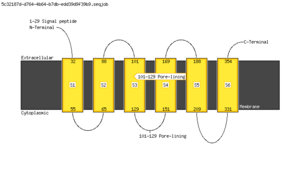
Memsat
|
<figure id="">[[File:|thumb|300px|<xr nolink id=""/> |
<figure id="hemo_memsat_P35462"></figure> | <figure id="hemo_memsat_P47863"></figure> | <figure id="hemo_memsat_Q9YDF8"></figure> |
Test the reference <xr id="Hemo_memsat_Q30201" />
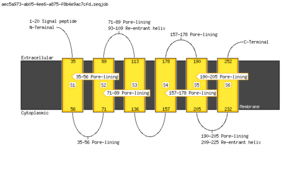
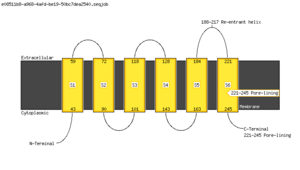
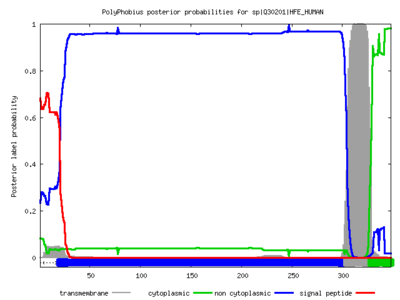
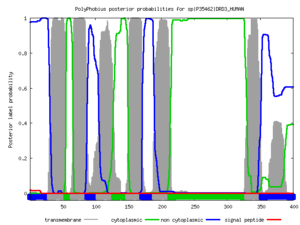
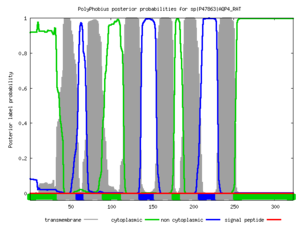
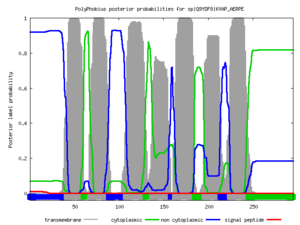
Polyphobius
| <figure id="hemo_poly_Q30201"></figure> | <figure id="hemo_poly_P35462"></figure> | <figure id="hemo_poly_P47863"></figure> | <figure id="hemo_poly_Q9YDF8"></figure> |
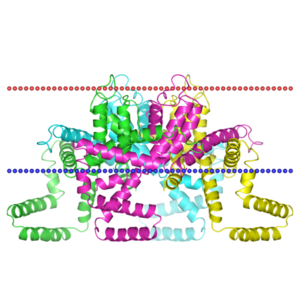
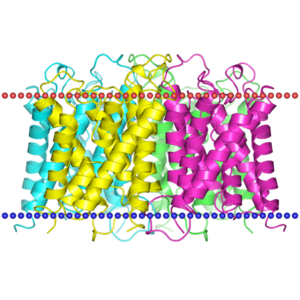
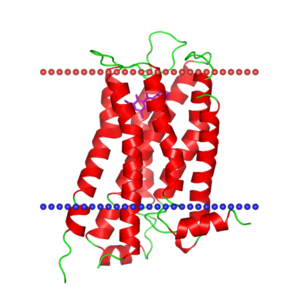
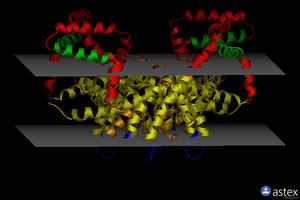
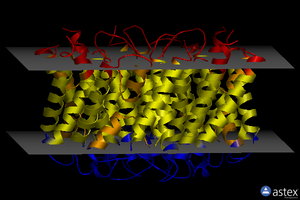
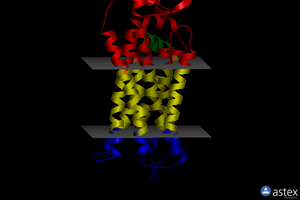
OPM
| <figure id="1orq_opm"></figure> | <figure id="2d57_opm"></figure> | <figure id="3pbl_opm"></figure> |
TMPDB
| <figure id="Hemo_1orq_lm"></figure> | <figure id="hemo_2d57_lm"></figure> | <figure id="hemo_3pbl_lm"></figure> |
Signal Peptides
GO Terms
References
<references />