Difference between revisions of "Task 5: Homology Modeling"
(→Single template) |
(→Single template) |
||
| Line 47: | Line 47: | ||
<figtable id="pymol str. al."> |
<figtable id="pymol str. al."> |
||
| − | {| class="wikitable" style="float: left; margin: 1em 0 0 0; border: 1px solid black;" cellpadding="0" |
+ | {| class="wikitable" style="float: left; margin: 1em 0 0 0; border: 1px solid black;" cellpadding="0"; align="center" |
| − | | align="center" | [[File:1qvo_std.png|thumb|200px| classical pairwise sequence alignment]] |
+ | | align="center" | [[File:1qvo_std.png|center|thumb|200px| classical pairwise sequence alignment]] |
| − | | align="center" | [[File:1qvo_2d.png|thumb|200px| inclusion of secondary structure information in the alignment ]] |
+ | | align="center" | [[File:1qvo_2d.png|center|thumb|200px| inclusion of secondary structure information in the alignment ]] |
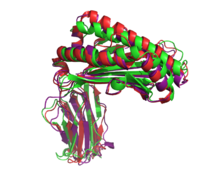
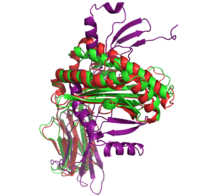
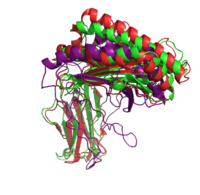
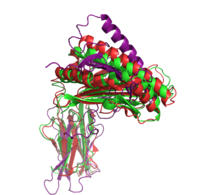
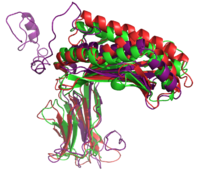
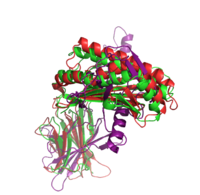
|+ style="caption-side: bottom; text-align: left" |<font size=1.5>'''Figure 1:'''Superposition of the target 1A6Z_A (green), the template 1QVO_A (red) and the model (purple). Two different alignment methods were used to create the input alignment. for Modeller. |
|+ style="caption-side: bottom; text-align: left" |<font size=1.5>'''Figure 1:'''Superposition of the target 1A6Z_A (green), the template 1QVO_A (red) and the model (purple). Two different alignment methods were used to create the input alignment. for Modeller. |
||
|} |
|} |
||
| Line 55: | Line 55: | ||
| + | |||
| − | rgehe |
||
<figtable id="pymol str. al."> |
<figtable id="pymol str. al."> |
||
| − | {| class="wikitable" style="float: left; margin: 1em 0 0 0; border: 1px solid black;" cellpadding="0" |
+ | {| class="wikitable" style="float: left; margin: 1em 0 0 0; border: 1px solid black;" cellpadding="0"; align="center" |
| align="center" | [[File:1s7x_std.png|thumb|200px| classical pairwise sequence alignment]] |
| align="center" | [[File:1s7x_std.png|thumb|200px| classical pairwise sequence alignment]] |
||
| align="center" | [[File:1s7x_2d.png|thumb|200px| inclusion of secondary structure information in the alignment ]] |
| align="center" | [[File:1s7x_2d.png|thumb|200px| inclusion of secondary structure information in the alignment ]] |
||
| Line 64: | Line 64: | ||
</figtable> |
</figtable> |
||
| + | |||
| − | sheth |
||
<figtable id="pymol str. al."> |
<figtable id="pymol str. al."> |
||
| − | {| class="wikitable" style="float: left; margin: 1em 0 0 0; border: 1px solid black;" cellpadding="0" |
+ | {| class="wikitable" style="float: left; margin: 1em 0 0 0; border: 1px solid black;" cellpadding="0"; align="center" |
| align="center" | [[File:1cd1_std.png|thumb|200px|classical pairwise sequence alignment ]] |
| align="center" | [[File:1cd1_std.png|thumb|200px|classical pairwise sequence alignment ]] |
||
| align="center" | [[File:1cd1_2d.png|thumb|200px|inclusion of secondary structure information in the alignment ]] |
| align="center" | [[File:1cd1_2d.png|thumb|200px|inclusion of secondary structure information in the alignment ]] |
||
Revision as of 23:27, 25 August 2013
1A6Z chain A was used as modeling target for all three methods.
Modeller
We used Modeller to create models based on a single template and multiple templates.
Single template
<css> table.colBasic2 { margin-left: auto; margin-right: auto; border: 2px solid black; border-collapse:collapse; width: 70%; } .colBasic2 th,td { padding: 3px; border: 2px solid black; } .colBasic2 td { text-align:left; } .colBasic2 tr th { background-color:#efefef; color: black;} .colBasic2 tr:first-child th { background-color:#adceff; color:black;} </css>
<figtable id="Modeller single">
| Template | Seq. identity | std Alignment | 2d alignment | curated Alignment | |||
|---|---|---|---|---|---|---|---|
| TM score | RMSD | TM score | RMSD | TM score | RMSD | ||
| 1QVO_A | 39% | 0.6241 | 3.647 | 0.5653 | 4.994 | ||
| 1S7X_A | 29% | 0.3355 | 15.806 | 0.2509 | 18.099 | ||
| 1CD1_A | 21% | 0.3640 | 18.066 | 0.4697 | 5.640 | ||
</figtable>
<xr id="Modeller single"/> lists the selected templates and the Modeller results for the different template structures and alignment methods. In addition to the standard pairwise sequence alignment based on dynamic programming, we also used Modeller's alignment.alig2dn() method to improve the alignment by including secondary structure information and improved the alignments manually. The RMSD and TM score are given for all models. The TM score ranges in the interval of (0, 1]. A value below 0.17 indicates a random similarity and a TM score above 0.5 corresponds to two structures with the same fold in CATH or SCOP (see TM score). Including the secondary structure information did only improve the model of the most distant homolog 1CD1_A.
<figtable id="pymol str. al.">
</figtable>
<figtable id="pymol str. al.">
</figtable>
<figtable id="pymol str. al.">
| "close" homology | |
| 1rjz_D | 0.34 |
| 1zag_A | 0.36 |
| 1QVO_A | 0.39 |
| distant homology: | |
| 3huj_C | 0.23 |
| 1cd1_A | 0.21 |
| 1VZY_A | 0.14 |
| MIX: |
| 1cd1_A 0.21 |
| 1QVO_A 0.39 |