Difference between revisions of "Canavan Task 2 - Sequence alignments"
(→Comparison of found sequences) |
(→Comparison of found sequences) |
||
| Line 120: | Line 120: | ||
|[[File:defaults.png|thumb|Default E-Values: as could be expected, the normal BLAST search is mostly contained in the PsiBLAST search with two iterations. HHBlits found a large number of different hits, with only 48 out of 274 common hits in common with the BLAST searches.]] |
|[[File:defaults.png|thumb|Default E-Values: as could be expected, the normal BLAST search is mostly contained in the PsiBLAST search with two iterations. HHBlits found a large number of different hits, with only 48 out of 274 common hits in common with the BLAST searches.]] |
||
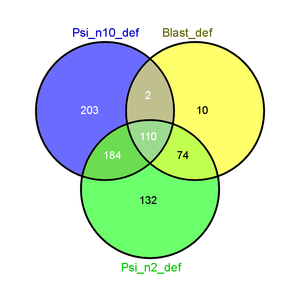
|[[File:Psi2_psi10_blast_defs.png|thumb|Taking PsiBLAST with 10 iterations into account brings in a large number of common sequences (110), which could be interesting since there seems to be high conversation among them.]] |
|[[File:Psi2_psi10_blast_defs.png|thumb|Taking PsiBLAST with 10 iterations into account brings in a large number of common sequences (110), which could be interesting since there seems to be high conversation among them.]] |
||
| − | |[[File:blast_psi2_e10_hhblits.png|thumb|Strict E-Value: ]] |
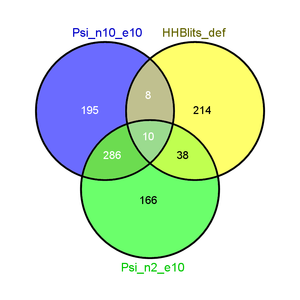
+ | |[[File:blast_psi2_e10_hhblits.png|thumb|Strict E-Values for PsiBLAST and default E-Value for HHBlits with 2 iterations: The number of common hits among all three is now substantially lower, while PsiBLAST with two and ten iterations share a great number of their hits. ]] |
|} |
|} |
||
Revision as of 14:53, 7 May 2012
Contents
Sequence
The native ASPA sequence that we used for the current task is shown below:
UniProt: P45381
>hsa:443 ASPA, ACY2, ASP; aspartoacylase; K01437 aspartoacylase [EC:3.5.1.15] (A)
MTSCHIAEEHIQKVAIFGGTHGNELTGVFLVKHWLENGAEIQRTGLEVKPFITNPRAVKK
CTRYIDCDLNRIFDLENLGKKMSEDLPYEVRRAQEINHLFGPKDSEDSYDIIFDLHNTTS
NMGCTLILEDSRNNFLIQMFHYIKTSLAPLPCYVYLIEHPSLKYATTRSIAKYPVGIEVG
PQPQGVLRADILDQMRKMIKHALDFIHHFNEGKEFPPCAIEVYKIIEKVDYPRDENGEIA
AIIHPNLQDQDWKPLHPGDPMFLTLDGKTIPLGGDCTVYPVFVNEAAYYEKKEAFAKTTK
LTLNAKSIRCCLH
Search
BLASTP
We ran BlastP on student machines with the big_80 as a reference database.
Command:
blastall -p blastp -d /mnt/project/pracstrucfunc12/data/big/big_80 -i P45381_wt.fasta -o blastp_p45381_wt_big80.out
| Parameters | default E-Value = 10 | E-Value 10e-10 |
| results | 196 | 94 |
| best E-Value | 1e-155 | 1e-155 |
| worst E-Value | 9.6 | e-15 |
| comment | Most of the resulting proteins are Aspartoacylases of other species. Most of the results with EValue > e-15 are Succinylglutamate Desuccinylases, which catalyze a reaction similar to Aspartoacylase. | The results are the same as for the first run, just with an earlier cutoff |
PSIBLAST
PSIBlast was used in the same fashion as BLAST, with the big_80 as the background database. Commands:
- Running 2 iterations and default E-Value 0.002
blastpgp -d /mnt/project/pracstrucfunc12/data/big/big_80 -i P45381_wt.fasta -o psiblast_it2_p45381_wt_big80.out -j 2
- 2 iterations, more strict E-value cutoff of 10E-10
blastpgp -d /mnt/project/pracstrucfunc12/data/big/big_80 -i P45381_wt.fasta -o psiblast_it2_h10e10_p45381_wt_big80.out -j 2 -h 10e-10
- 10 iterations, default Evalue 0.002
blastpgp -d /mnt/project/pracstrucfunc12/data/big/big_80 -i P45381_wt.fasta -o psiblast_it10_p45381_wt_big80.out -j 10
- 10 iterations, E-value cutoff 10E-10
blastpgp -d /mnt/project/pracstrucfunc12/data/big/big_80 -i P45381_wt.fasta -o psiblast_it10_h10e10_p45381_wt_big80.out -j 10 -h 10e-10
| Parameters | it2, def E-Value (0.002) | it2 E-Value 10e-10 | it10 def E-Value (0.002) | it10 E-Value 10e-10 |
| time | ~2m30 | ~2m30 | ~10m | time: ~10m |
| results | 500 | 93 | 500 | 500 |
| best E-Value | 1e-142 | 1e-145 | 5e-70 | 7e-70 |
| worst E-Value | 3e-4 | 2e-29 | 8e-38 | 1e-38 |
| comments | Results with best EValues are mostly Aspartoacylases, Sequences previously not found are mostly Succinylglutamate Desuccinylases | results mainly Aspartoacylases | - converged after 8 rounds - most significant results include more Succinylglutamate Desuccinylases than Aspartoacylases | - all 10 iterations were done (no early convergence) - aspartoacylases slightly more frequent in lower E-Values (< E-58), but no significant difference in E-Values for aspas and succis |
HHBLITS
Run HHBlits on student machines with Uniprot20 database.
Commands
- 2 iterations:
hhblits -i P45381_wt.fasta -d /mnt/project/pracstrucfunc12/data/hhblits/uniprot20_current -o hhblits_p45381_def.out
- 8 iterations:
hhblits -i P45381_wt.fasta -d /mnt/project/pracstrucfunc12/data/hhblits/uniprot20_current -n 8 -o hhblits_p45381_n10.out
-n number of iterations (def 2)
Summary and Comparison
Along with the expactations one can find more hits with Psi-Blast than with a simple Blast search.
In general, one can distinguish between two kinds of proteins, that frequently are identified by the sequence searches:
- Aspartoacylases
- Succinylglutamate Desuccinylases
BlastP
A simple blast search yields only about 90 significant hits it one considers a threshold of 10e-10 as a significance cutoff. As one can see in Figure ??, the restriction of the Evalue results in less hits with a low sequence similarity.
Psi Blast
Increasing the amount of iterations performed in a PSI-Blast search, obviously increases the running time. One can see, that the best ranked hits have lower E-Values than the best Hits of the runs with less iterations. Yet, there are more hits found with better E-Values, which is not surprising because more homologues of significant profile sequences will be found.
When restricting the E-Value Cutoff for the profile built-up, we found that more hits are classified as Aspartoacylases than as Succinylglutamate Desuccinylases. The running time, as well as the E-Values of the resulting hits did not change significantly.
HHBlits
Comparison of found sequences
| Parameters | it 2 | it 8 |
| time | 2m50 | ~6m |
| results | 274 | 500 |
| best E-Value | 2e-110 | 2.9e-68 |
| worst E-Value | 0.0011 | 9.5e-09 |
| comment | mixed results with Aspartoacylases and Succi | very varying results: Aspartoacylasen, Succinylasen, Zinc Proteins |