Difference between revisions of "Sequence Alignments HEXA"
(→Multiple Alignments) |
(→Sequence Alignments) |
||
| Line 14: | Line 14: | ||
* HHSearch |
* HHSearch |
||
For the HHSearch tool we used the online server for [http://toolkit.tuebingen.mpg.de/hhpred HHSearch]. |
For the HHSearch tool we used the online server for [http://toolkit.tuebingen.mpg.de/hhpred HHSearch]. |
||
| + | |||
| + | |||
| + | '''Result Statistics''' |
||
| + | |||
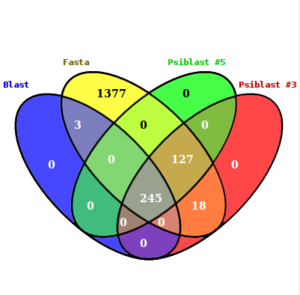
| + | [[Image:comparison_alignments.png|thumb|center|Overlap between the different Alignments Methods.)]] |
||
== Multiple Alignments == |
== Multiple Alignments == |
||
Revision as of 10:58, 23 May 2011
Sequence Alignments
Sequence Searches:
- FASTA
/bin/fasta36 seq.fasta /data/blast/nr/nr > fasta_out.txt
- BLAST
blastall -p blastp -d /data/blast/nr/nr -i mult_seq.fasta > blast_out.txt
- PSIBLAST
blastpgn -i seq.fasta -j <#iterations> -h <e-value threshold> -d /data/blast/nr/nr > psiblast_out.txt
- HHSearch
For the HHSearch tool we used the online server for HHSearch.
Result Statistics
Multiple Alignments
- Cobalt
Download Cobalt from ftp://ftp.ncbi.nlm.nih.gov/pub/cobalt/executables/2.0.1/ (ncbi-cobalt-2.0.1-x64-linux.tar). Uncompress the archive file with tar xfz ncbi-cobalt-2.0.1-x64-linux.tar and change directory to the uncompressed cobalt directoy. Call: ./cobalt -i mult_seq.fasta -norps T > cobalt_out.aln
- ClustalW
clustalw -infile=mult_seq.fasta > clustalW_out.aln
- Muscle
muscle -in mult_seq.fasta -out muscle_out.aln -clw
- T-Coffee
t_coffee -seq mult_seq.fasta
- T-Coffee (3D)
t_coffee -seq mult_seq.fasta -mode expresso